You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000555_01764
You are here: Home > Sequence: MGYG000000555_01764
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
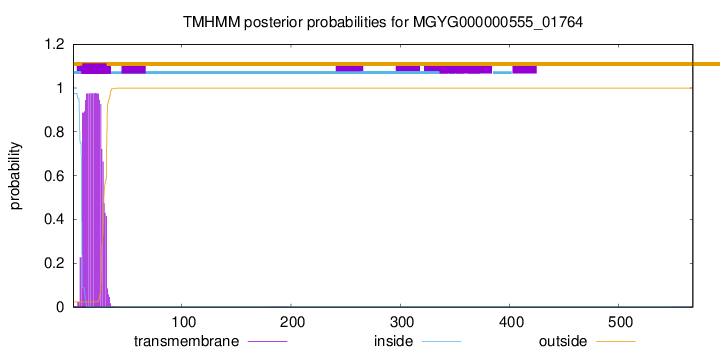
TMHMM annotations
Basic Information help
| Species | Cellulosilyticum sp900556665 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Cellulosilyticaceae; Cellulosilyticum; Cellulosilyticum sp900556665 | |||||||||||
| CAZyme ID | MGYG000000555_01764 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1077; End: 2783 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 61 | 277 | 1.7e-55 | 0.973568281938326 |
| CBM2 | 475 | 557 | 4.9e-22 | 0.8217821782178217 |
| CBM13 | 310 | 459 | 4.9e-19 | 0.7446808510638298 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2382 | Fes | 1.69e-41 | 28 | 278 | 44 | 295 | Enterochelin esterase or related enzyme [Inorganic ion transport and metabolism]. |
| pfam00756 | Esterase | 9.52e-18 | 61 | 271 | 1 | 239 | Putative esterase. This family contains Esterase D. However it is not clear if all members of the family have the same function. This family is related to the pfam00135 family. |
| smart00637 | CBD_II | 5.29e-17 | 480 | 558 | 1 | 80 | CBD_II domain. |
| pfam00553 | CBM_2 | 1.32e-15 | 479 | 559 | 7 | 87 | Cellulose binding domain. Two tryptophan residues are involved in cellulose binding. Cellulose binding domain found in bacteria. |
| pfam14200 | RicinB_lectin_2 | 1.82e-14 | 346 | 436 | 1 | 89 | Ricin-type beta-trefoil lectin domain-like. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACZ98598.1 | 0.0 | 1 | 568 | 1 | 566 |
| ADZ84408.1 | 6.24e-293 | 3 | 568 | 6 | 569 |
| ACZ98596.1 | 1.80e-131 | 290 | 568 | 395 | 667 |
| QNU68935.1 | 2.87e-118 | 29 | 337 | 18 | 314 |
| AGF55271.1 | 6.00e-118 | 6 | 286 | 2 | 286 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1JJF_A | 4.66e-116 | 25 | 284 | 3 | 263 | ChainA, Endo-1,4-beta-xylanase Z [Acetivibrio thermocellus] |
| 1JT2_A | 1.32e-115 | 25 | 284 | 3 | 263 | ChainA, PROTEIN (ENDO-1,4-BETA-XYLANASE Z) [Acetivibrio thermocellus] |
| 7B5V_A | 2.86e-52 | 38 | 271 | 121 | 377 | ChainA, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B5V_B Chain B, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B6B_A Chain A, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B6B_B Chain B, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836] |
| 6MOT_A | 5.52e-45 | 38 | 267 | 99 | 346 | ChainA, Isoamylase N-terminal domain protein [Bacteroides intestinalis DSM 17393] |
| 6MOU_A | 9.14e-45 | 38 | 267 | 120 | 367 | ChainA, Isoamylase N-terminal domain protein [Bacteroides intestinalis DSM 17393],6MOU_B Chain B, Isoamylase N-terminal domain protein [Bacteroides intestinalis DSM 17393] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P10478 | 3.08e-110 | 7 | 455 | 3 | 425 | Endo-1,4-beta-xylanase Z OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=xynZ PE=1 SV=3 |
| D5EXZ4 | 2.03e-29 | 43 | 271 | 420 | 661 | Carbohydrate acetyl esterase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=axe1-6A PE=1 SV=1 |
| D5EY13 | 1.03e-27 | 47 | 278 | 493 | 724 | Endo-1,4-beta-xylanase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=xyn10D-fae1A PE=1 SV=1 |
| P31471 | 9.04e-26 | 37 | 270 | 126 | 379 | Uncharacterized protein YieL OS=Escherichia coli (strain K12) OX=83333 GN=yieL PE=4 SV=3 |
| P51584 | 1.26e-16 | 67 | 230 | 840 | 1016 | Endo-1,4-beta-xylanase Y OS=Acetivibrio thermocellus OX=1515 GN=xynY PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
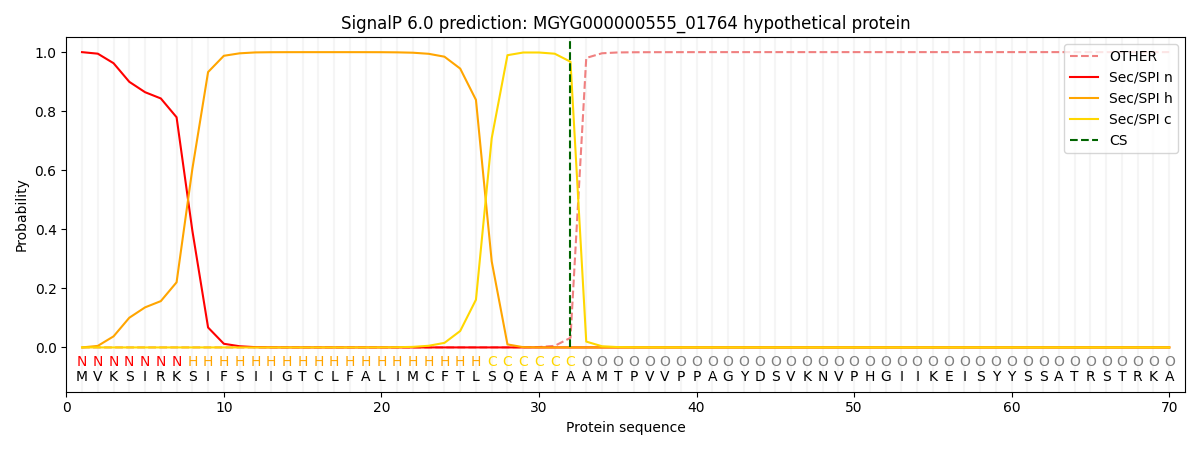
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000551 | 0.998582 | 0.000325 | 0.000206 | 0.000156 | 0.000148 |