You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000559_02414
You are here: Home > Sequence: MGYG000000559_02414
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paraprevotella sp900546665 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Paraprevotella; Paraprevotella sp900546665 | |||||||||||
| CAZyme ID | MGYG000000559_02414 | |||||||||||
| CAZy Family | GH30 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5957; End: 10183 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH30 | 55 | 504 | 5e-125 | 0.9856115107913669 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02055 | Glyco_hydro_30 | 7.40e-26 | 64 | 443 | 2 | 348 | Glycosyl hydrolase family 30 TIM-barrel domain. |
| COG5520 | XynC | 7.03e-20 | 147 | 510 | 96 | 432 | O-Glycosyl hydrolase [Cell wall/membrane/envelope biogenesis]. |
| pfam17189 | Glyco_hydro_30C | 1.15e-04 | 450 | 504 | 7 | 63 | Glycosyl hydrolase family 30 beta sandwich domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCD34402.1 | 9.52e-110 | 7 | 505 | 6 | 501 |
| QIU97167.1 | 3.74e-108 | 27 | 505 | 31 | 493 |
| QDO68217.1 | 6.72e-108 | 27 | 504 | 30 | 490 |
| ALJ59961.1 | 1.72e-107 | 27 | 504 | 29 | 489 |
| ABR38208.1 | 3.23e-107 | 27 | 504 | 28 | 499 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2WNW_A | 1.44e-16 | 66 | 505 | 48 | 441 | Thecrystal structure of SrfJ from salmonella typhimurium [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],2WNW_B The crystal structure of SrfJ from salmonella typhimurium [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O16580 | 2.85e-23 | 50 | 478 | 85 | 484 | Putative glucosylceramidase 1 OS=Caenorhabditis elegans OX=6239 GN=gba-1 PE=1 SV=2 |
| O16581 | 6.37e-23 | 42 | 476 | 74 | 480 | Putative glucosylceramidase 2 OS=Caenorhabditis elegans OX=6239 GN=gba-2 PE=3 SV=2 |
| Q9UB00 | 2.35e-13 | 66 | 476 | 102 | 484 | Putative glucosylceramidase 4 OS=Caenorhabditis elegans OX=6239 GN=gba-4 PE=3 SV=2 |
| Q7M4T0 | 2.01e-09 | 50 | 357 | 55 | 321 | Endo-1,6-beta-D-glucanase OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=neg-1 PE=1 SV=2 |
| Q4WBR2 | 8.96e-07 | 40 | 506 | 43 | 485 | Endo-1,6-beta-D-glucanase neg1 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=neg1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
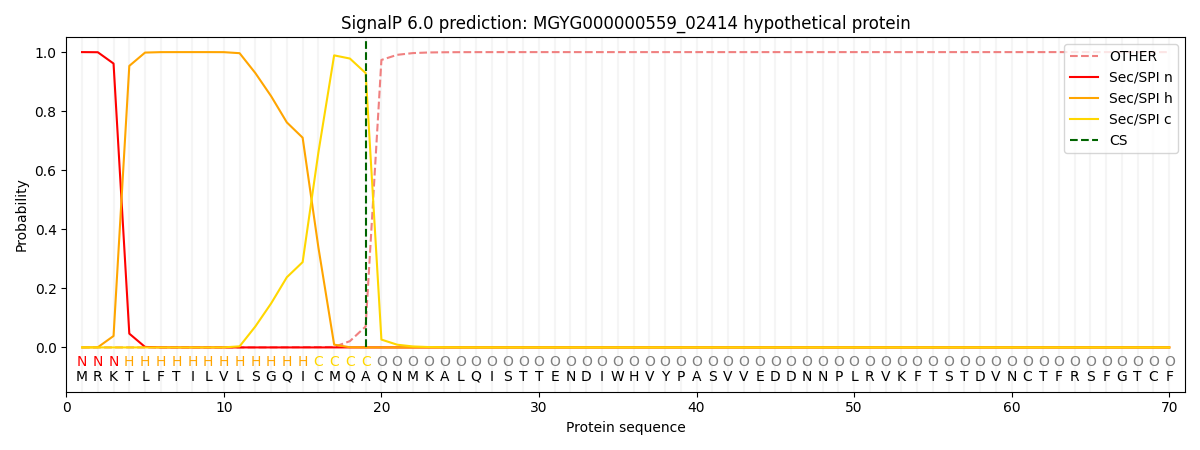
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000268 | 0.999073 | 0.000151 | 0.000180 | 0.000155 | 0.000147 |
