You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000575_02667
You are here: Home > Sequence: MGYG000000575_02667
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA11452 sp003526375 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; UBA1829; UBA11452; UBA11452 sp003526375 | |||||||||||
| CAZyme ID | MGYG000000575_02667 | |||||||||||
| CAZy Family | GH89 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3555; End: 5888 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH89 | 343 | 653 | 3.1e-23 | 0.43891402714932126 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam03648 | Glyco_hydro_67N | 0.006 | 14 | 107 | 34 | 120 | Glycosyl hydrolase family 67 N-terminus. Alpha-glucuronidases, components of an ensemble of enzymes central to the recycling of photosynthetic biomass, remove the alpha-1,2 linked 4-O-methyl glucuronic acid from xylans. This family represents the N-terminal region of alpha-glucuronidase. The N-terminal domain forms a two-layer sandwich, each layer being formed by a beta sheet of five strands. A further two helices form part of the interface with the central, catalytic, module (pfam07488). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBL05600.1 | 1.74e-215 | 2 | 745 | 26 | 758 |
| EAQ48518.1 | 1.41e-07 | 8 | 167 | 49 | 201 |
| AQT67390.1 | 4.53e-07 | 76 | 267 | 106 | 277 |
| ASU35870.1 | 8.94e-07 | 49 | 512 | 6 | 473 |
| QGY77385.1 | 9.32e-07 | 90 | 517 | 92 | 550 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
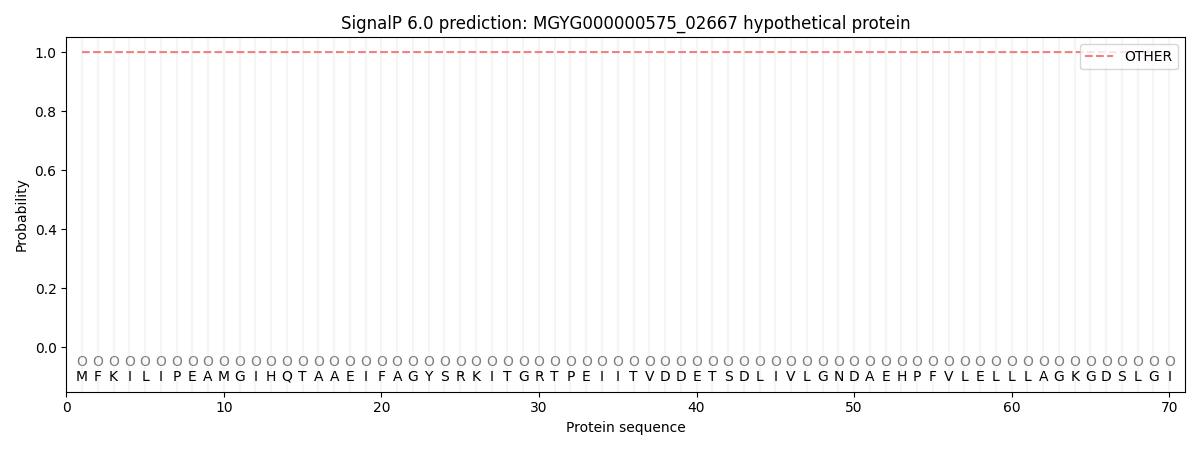
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000034 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
