You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000592_01295
You are here: Home > Sequence: MGYG000000592_01295
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; UBA932; RC9; | |||||||||||
| CAZyme ID | MGYG000000592_01295 | |||||||||||
| CAZy Family | PL35 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4441; End: 6102 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL35 | 318 | 495 | 3.6e-50 | 0.9553072625698324 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07940 | Hepar_II_III | 2.07e-12 | 323 | 511 | 29 | 208 | Heparinase II/III-like protein. This family features sequences that are similar to a region of the Flavobacterium heparinum proteins heparinase II and heparinase III. The former is known to degrade heparin and heparin sulphate, whereas the latter predominantly degrades heparin sulphate. Both are secreted into the periplasmic space upon induction with heparin. |
| pfam05147 | LANC_like | 0.008 | 28 | 180 | 15 | 162 | Lanthionine synthetase C-like protein. Lanthionines are thioether bridges that are putatively generated by dehydration of Ser and Thr residues followed by addition of cysteine residues within the peptide. This family contains the lanthionine synthetase C-like proteins 1 and 2 which are related to the bacterial lanthionine synthetase components C (LanC). LANCL1 (P40 seven-transmembrane-domain protein) and LANCL2 (testes-specific adriamycin sensitivity protein) are thought to be peptide-modifying enzyme components in eukaryotic cells. Both proteins are produced in large quantities in the brain and testes and may have role in the immune surveillance of these organs. Lanthionines are found in lantibiotics, which are peptide-derived, post-translationally modified antimicrobials produced by several bacterial strains. This region contains seven internal repeats. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QGT72631.1 | 7.54e-191 | 1 | 550 | 98 | 640 |
| SCV07259.1 | 1.07e-190 | 1 | 550 | 98 | 640 |
| QNL40060.1 | 1.07e-190 | 1 | 550 | 98 | 640 |
| QRQ54740.1 | 1.07e-190 | 1 | 550 | 98 | 640 |
| ALJ47854.1 | 1.07e-190 | 1 | 550 | 98 | 640 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3A0O_A | 1.38e-10 | 72 | 493 | 259 | 681 | Crystalstructure of alginate lyase from Agrobacterium tumefaciens C58 [Agrobacterium fabrum str. C58],3A0O_B Crystal structure of alginate lyase from Agrobacterium tumefaciens C58 [Agrobacterium fabrum str. C58] |
| 3AFL_A | 3.17e-10 | 72 | 248 | 259 | 435 | Crystalstructure of exotype alginate lyase Atu3025 H531A complexed with alginate trisaccharide [Agrobacterium fabrum str. C58] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
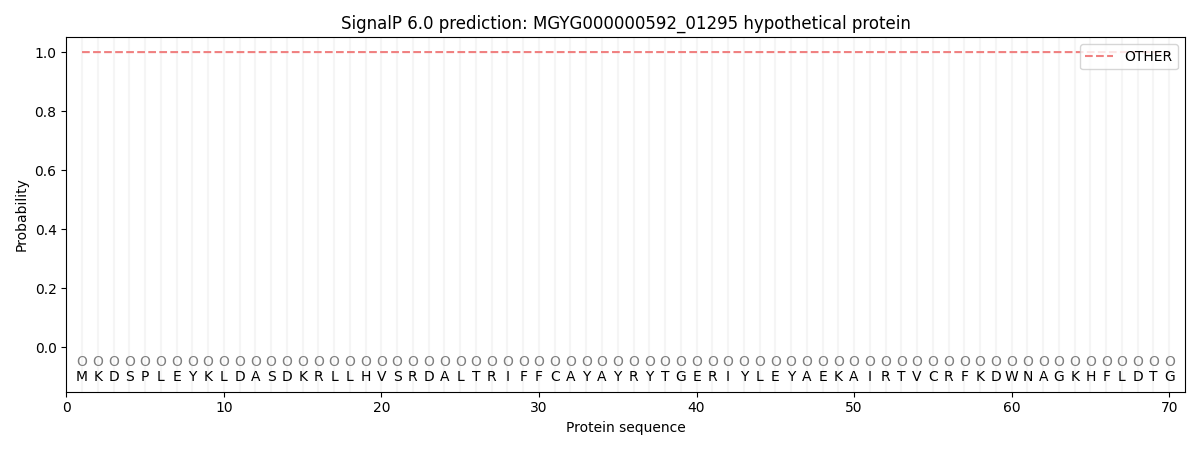
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000058 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
