You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000600_01411
You are here: Home > Sequence: MGYG000000600_01411
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-791 sp000431495 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; CAG-791; CAG-791 sp000431495 | |||||||||||
| CAZyme ID | MGYG000000600_01411 | |||||||||||
| CAZy Family | GH8 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 27958; End: 30225 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3405 | BcsZ | 8.01e-73 | 411 | 755 | 37 | 351 | Endo-1,4-beta-D-glucanase Y [Carbohydrate transport and metabolism]. |
| cd07712 | MBLAC2-like_MBL-fold | 4.17e-39 | 72 | 254 | 10 | 182 | uncharacterized human metallo-beta-lactamase domain-containing protein 2 and related proteins; MBL-fold metallo hydrolase domain. Includes the MBL-fold metallo hydrolase domain of uncharacterized human MBLAC2 and related proteins. Members of this subgroup belong to the MBL-fold metallo-hydrolase superfamily which is comprised mainly of hydrolytic enzymes which carry out a variety of biological functions. |
| cd06262 | metallo-hydrolase-like_MBL-fold | 1.99e-20 | 67 | 214 | 6 | 147 | mainly hydrolytic enzymes and related proteins which carry out various biological functions; MBL-fold metallohydrolase domain. Members of the MBL-fold metallohydrolase superfamily are mainly hydrolytic enzymes which carry out a variety of biological functions. The class B metal beta-lactamases (MBLs) for which this fold was named perform only a small fraction of the activities included in this superfamily. Activities carried out by superfamily members include class B beta-lactamases which can catalyze the hydrolysis of a wide range of beta-lactam antibiotics, hydroxyacylglutathione hydrolases (also called glyoxalase II) which hydrolyze S-d-lactoylglutathione to d-lactate in the second step of the glycoxlase system, AHL lactonases which catalyze the hydrolysis and opening of the homoserine lactone rings of acyl homoserine lactones (AHLs), persulfide dioxygenase which catalyze the oxidation of glutathione persulfide to glutathione and persulfite in the mitochondria, flavodiiron proteins which catalyze the reduction of oxygen and/or nitric oxide to water or nitrous oxide respectively, cleavage and polyadenylation specificity factors such as the Int9 and Int11 subunits of Integrator, Sdsa1-like and AtsA-like arylsulfatases, 5'-exonucleases human SNM1A and yeast Pso2p, ribonuclease J which has both 5'-3' exoribonucleolytic and endonucleolytic activity and ribonuclease Z which catalyzes the endonucleolytic removal of the 3' extension of the majority of tRNA precursors, cyclic nucleotide phosphodiesterases which decompose cyclic adenosine and guanosine 3', 5'-monophosphate (cAMP and cGMP) respectively, insecticide hydrolases, and proteins required for natural transformation competence. The diversity of biological roles is reflected in variations in the active site metallo-chemistry, for example classical members of the superfamily are di-, or less commonly mono-, zinc-ion-dependent hydrolases, human persulfide dioxygenase ETHE1 is a mono-iron binding member of the superfamily; Arabidopsis thaliana hydroxyacylglutathione hydrolases incorporates iron, manganese, and zinc in its dinuclear metal binding site, and flavodiiron proteins contains a diiron site. |
| smart00849 | Lactamase_B | 4.69e-20 | 72 | 215 | 1 | 137 | Metallo-beta-lactamase superfamily. Apart from the beta-lactamases a number of other proteins contain this domain. These proteins include thiolesterases, members of the glyoxalase II family, that catalyse the hydrolysis of S-D-lactoyl-glutathione to form glutathione and D-lactic acid and a competence protein that is essential for natural transformation in Neisseria gonorrhoeae and could be a transporter involved in DNA uptake. Except for the competence protein these proteins bind two zinc ions per molecule as cofactor. |
| cd07729 | AHL_lactonase_MBL-fold | 1.74e-18 | 72 | 254 | 33 | 236 | quorum-quenching N-acyl-homoserine lactonase, MBL-fold metallo-hydrolase domain. Acyl Homoserine Lactones (also known as AHLs) are signal molecules which coordinate gene expression in quorum sensing, in many Gram-negative bacteria. Quorum-quenching N-acyl-homoserine lactonase (also known as AHL lactonase, N-acyl-L-homoserine lactone hydrolase, EC 3.1.1.81) catalyzes the hydrolysis and opening of the homoserine lactone rings of AHLs, a reaction that can block quorum sensing. These enzymes belong to the MBL-fold metallo-hydrolase superfamily which is comprised mainly of hydrolytic enzymes which carry out a variety of biological functions. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNO17280.1 | 1.13e-165 | 388 | 754 | 26 | 380 |
| CBL10560.1 | 2.27e-160 | 388 | 755 | 27 | 383 |
| CBL13086.1 | 2.27e-160 | 388 | 755 | 27 | 383 |
| EEU99943.1 | 1.16e-157 | 388 | 755 | 27 | 383 |
| VCV21314.1 | 1.16e-157 | 388 | 755 | 27 | 383 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5YXT_A | 3.10e-141 | 385 | 754 | 26 | 375 | Glycosidehydrolase family 8 Xylanase [Paenibacillus barengoltzii G22],5YXT_B Glycoside hydrolase family 8 Xylanase [Paenibacillus barengoltzii G22],5YXT_C Glycoside hydrolase family 8 Xylanase [Paenibacillus barengoltzii G22],5YXT_D Glycoside hydrolase family 8 Xylanase [Paenibacillus barengoltzii G22] |
| 1WU4_A | 7.86e-137 | 388 | 754 | 30 | 378 | ChainA, xylanase Y [Halalkalibacterium halodurans C-125],1WU5_A Chain A, xylanase Y [Halalkalibacterium halodurans C-125] |
| 3A3V_A | 3.12e-136 | 388 | 754 | 30 | 378 | ChainA, Xylanase Y [Halalkalibacterium halodurans] |
| 2DRR_A | 4.41e-136 | 388 | 754 | 30 | 378 | ChainA, Xylanase Y [Halalkalibacterium halodurans C-125] |
| 1WU6_A | 6.22e-136 | 388 | 754 | 30 | 378 | ChainA, xylanase Y [Halalkalibacterium halodurans C-125] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9KB30 | 3.28e-136 | 388 | 754 | 30 | 378 | Reducing end xylose-releasing exo-oligoxylanase OS=Alkalihalobacillus halodurans (strain ATCC BAA-125 / DSM 18197 / FERM 7344 / JCM 9153 / C-125) OX=272558 GN=BH2105 PE=1 SV=1 |
| A1A048 | 5.20e-129 | 385 | 753 | 25 | 377 | Reducing end xylose-releasing exo-oligoxylanase OS=Bifidobacterium adolescentis (strain ATCC 15703 / DSM 20083 / NCTC 11814 / E194a) OX=367928 GN=xylA PE=1 SV=1 |
| A0A0S2UQQ5 | 7.60e-129 | 389 | 755 | 31 | 380 | Reducing-end xylose-releasing exo-oligoxylanase Rex8A OS=Paenibacillus barcinonensis OX=198119 GN=rex8A PE=1 SV=1 |
| P37699 | 2.74e-18 | 392 | 752 | 68 | 389 | Endoglucanase C OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCC PE=1 SV=2 |
| P37701 | 3.65e-18 | 392 | 752 | 68 | 389 | Endoglucanase 2 OS=Ruminiclostridium josui OX=1499 GN=celB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
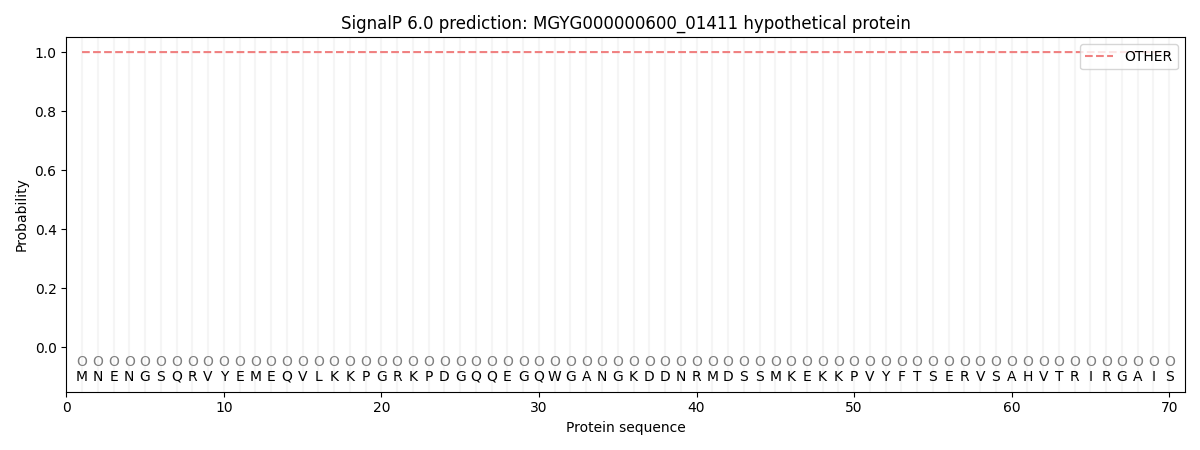
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000052 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
