You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000609_00110
You are here: Home > Sequence: MGYG000000609_00110
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; TANB77; UBA1234; UBA1234; | |||||||||||
| CAZyme ID | MGYG000000609_00110 | |||||||||||
| CAZy Family | GT0 | |||||||||||
| CAZyme Description | UDP-N-acetylglucosamine 2-epimerase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2848; End: 5070 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| TIGR00236 | wecB | 5.90e-166 | 364 | 730 | 1 | 365 | UDP-N-acetylglucosamine 2-epimerase. This cytosolic enzyme converts UDP-N-acetyl-D-glucosamine to UDP-N-acetyl-D-mannosamine. In E. coli, this is the first step in the pathway of enterobacterial common antigen biosynthesis.Members of this orthology group have many gene symbols, often reflecting the overall activity of the pathway and/or operon that includes it. Symbols include epsC (exopolysaccharide C) in Burkholderia solanacerum, cap8P (type 8 capsule P) in Staphylococcus aureus, and nfrC in an older designation based on the effects of deletion on phage N4 adsorption. Epimerase activity was also demonstrated in a bifunctional rat enzyme, for which the N-terminal domain appears to be orthologous. The set of proteins found above the suggested cutoff includes E. coli WecB in one of two deeply branched clusters and the rat UDP-N-acetylglucosamine 2-epimerase domain in the other. [Cell envelope, Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides] |
| COG0381 | WecB | 1.20e-164 | 364 | 737 | 4 | 379 | UDP-N-acetylglucosamine 2-epimerase [Cell wall/membrane/envelope biogenesis]. |
| cd03786 | GTB_UDP-GlcNAc_2-Epimerase | 4.31e-135 | 365 | 727 | 1 | 365 | UDP-N-acetylglucosamine 2-epimerase and similar proteins. Bacterial members of the UDP-N-Acetylglucosamine (GlcNAc) 2-Epimerase family (EC 5.1.3.14) are known to catalyze the reversible interconversion of UDP-GlcNAc and UDP-N-acetylmannosamine (UDP-ManNAc). The enzyme serves to produce an activated form of ManNAc residues (UDP-ManNAc) for use in the biosynthesis of a variety of cell surface polysaccharides; The mammalian enzyme is bifunctional, catalyzing both the inversion of stereochemistry at C-2 and the hydrolysis of the UDP-sugar linkage to generate free ManNAc. It also catalyzes the phosphorylation of ManNAc to generate ManNAc 6-phosphate, a precursor to salic acids. In mammals, sialic acids are found at the termini of oligosaccharides in a large variety of cell surface glycoconjugates and are key mediators of cell-cell recognition events. Mutations in human members of this family have been associated with Sialuria, a rare disease caused by the disorders of sialic acid metabolism. This family belongs to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. |
| pfam02350 | Epimerase_2 | 1.02e-130 | 386 | 727 | 3 | 336 | UDP-N-acetylglucosamine 2-epimerase. This family consists of UDP-N-acetylglucosamine 2-epimerases EC:5.1.3.14 this enzyme catalyzes the production of UDP-ManNAc from UDP-GlcNAc. Note that some of the enzymes is this family are bifunctional, in these instances Pfam matches only the N-terminal half of the protein suggesting that the additional C-terminal part (when compared to mono-functional members of this family) is responsible for the UPD-N-acetylmannosamine kinase activity of these enzymes. This hypothesis is further supported by the assumption that the C-terminal part of rat Gne is the kinase domain. |
| cd06853 | GT_WecA_like | 9.25e-106 | 40 | 290 | 1 | 249 | This subfamily contains Escherichia coli WecA, Bacillus subtilis TagO and related proteins. WecA is an UDP-N-acetylglucosamine (GlcNAc):undecaprenyl-phosphate (Und-P) GlcNAc-1-phosphate transferase that catalyzes the formation of a phosphodiester bond between a membrane-associated undecaprenyl-phosphate molecule and N-acetylglucosamine 1-phosphate, which is usually donated by a soluble UDP-N-acetylglucosamine precursor. WecA participates in the biosynthesis of O antigen LPS in many enteric bacteria and is also involved in the biosynthesis of enterobacterial common antigen. A conserved short sequence motif and a conserved arginine at a cytosolic loop of this integral membrane protein were shown to be critical in recognition of substrate UDP-N-acetylglucosamine. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOR22598.1 | 5.18e-166 | 365 | 736 | 5 | 376 |
| QES71705.1 | 4.72e-164 | 365 | 736 | 5 | 376 |
| ALB48231.2 | 4.72e-164 | 365 | 736 | 5 | 376 |
| QGH21225.1 | 9.45e-164 | 365 | 730 | 5 | 370 |
| ALP89282.1 | 9.45e-164 | 365 | 730 | 5 | 370 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3BEO_A | 9.16e-158 | 361 | 730 | 6 | 373 | AStructural Basis for the allosteric regulation of non-hydrolyzing UDP-GlcNAc 2-epimerases [Bacillus anthracis],3BEO_B A Structural Basis for the allosteric regulation of non-hydrolyzing UDP-GlcNAc 2-epimerases [Bacillus anthracis] |
| 4FKZ_A | 1.50e-151 | 363 | 737 | 3 | 374 | Crystalstructure of Bacillus subtilis UDP-GlcNAc 2-epimerase in complex with UDP-GlcNAc and UDP [Bacillus subtilis subsp. subtilis str. 168],4FKZ_B Crystal structure of Bacillus subtilis UDP-GlcNAc 2-epimerase in complex with UDP-GlcNAc and UDP [Bacillus subtilis subsp. subtilis str. 168] |
| 3OT5_A | 1.81e-148 | 363 | 739 | 27 | 400 | 2.2Angstrom Resolution Crystal Structure of putative UDP-N-acetylglucosamine 2-epimerase from Listeria monocytogenes [Listeria monocytogenes EGD-e],3OT5_B 2.2 Angstrom Resolution Crystal Structure of putative UDP-N-acetylglucosamine 2-epimerase from Listeria monocytogenes [Listeria monocytogenes EGD-e],3OT5_C 2.2 Angstrom Resolution Crystal Structure of putative UDP-N-acetylglucosamine 2-epimerase from Listeria monocytogenes [Listeria monocytogenes EGD-e],3OT5_D 2.2 Angstrom Resolution Crystal Structure of putative UDP-N-acetylglucosamine 2-epimerase from Listeria monocytogenes [Listeria monocytogenes EGD-e] |
| 1O6C_A | 1.54e-145 | 363 | 737 | 3 | 374 | Crystalstructure of UDP-N-acetylglucosamine 2-epimerase [Bacillus subtilis],1O6C_B Crystal structure of UDP-N-acetylglucosamine 2-epimerase [Bacillus subtilis] |
| 5ENZ_A | 4.41e-144 | 365 | 739 | 3 | 374 | S.aureus MnaA-UDP co-structure [Staphylococcus aureus],5ENZ_B S. aureus MnaA-UDP co-structure [Staphylococcus aureus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P45360 | 5.07e-168 | 363 | 739 | 3 | 380 | Putative UDP-N-acetylglucosamine 2-epimerase OS=Clostridium acetobutylicum (strain ATCC 824 / DSM 792 / JCM 1419 / LMG 5710 / VKM B-1787) OX=272562 GN=CA_C2874 PE=3 SV=2 |
| Q9X0C4 | 4.11e-151 | 364 | 733 | 2 | 371 | Putative UDP-N-acetylglucosamine 2-epimerase OS=Thermotoga maritima (strain ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8) OX=243274 GN=TM_1034 PE=3 SV=1 |
| P39131 | 6.24e-151 | 363 | 737 | 3 | 374 | UDP-N-acetylglucosamine 2-epimerase OS=Bacillus subtilis (strain 168) OX=224308 GN=mnaA PE=1 SV=1 |
| Q9L6R5 | 2.24e-135 | 364 | 729 | 1 | 372 | UDP-N-acetylglucosamine 2-epimerase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=wecB PE=3 SV=1 |
| Q8XAR8 | 2.24e-135 | 364 | 727 | 1 | 370 | UDP-N-acetylglucosamine 2-epimerase OS=Escherichia coli O157:H7 OX=83334 GN=wecB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000044 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
TMHMM Annotations download full data without filtering help
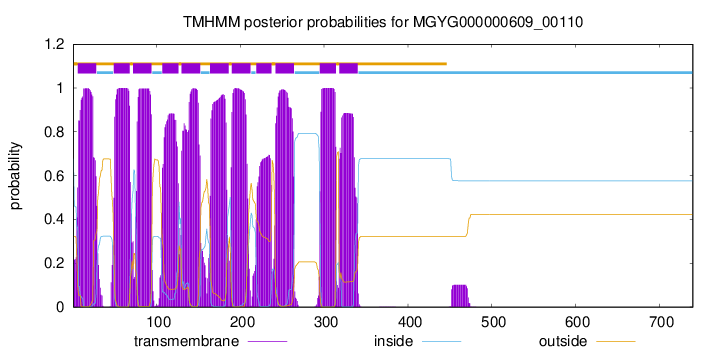
| start | end |
|---|---|
| 6 | 28 |
| 49 | 68 |
| 72 | 94 |
| 107 | 126 |
| 130 | 152 |
| 164 | 186 |
| 190 | 212 |
| 219 | 237 |
| 242 | 264 |
| 295 | 314 |
| 318 | 340 |
