You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000633_00009
You are here: Home > Sequence: MGYG000000633_00009
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Victivallis sp900768515 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; Victivallaceae; Victivallis; Victivallis sp900768515 | |||||||||||
| CAZyme ID | MGYG000000633_00009 | |||||||||||
| CAZy Family | GT19 | |||||||||||
| CAZyme Description | Lipid-A-disaccharide synthase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 11617; End: 12774 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT19 | 5 | 362 | 4.2e-89 | 0.9858757062146892 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK00025 | lpxB | 1.33e-91 | 5 | 364 | 4 | 359 | lipid-A-disaccharide synthase; Reviewed |
| COG0763 | LpxB | 3.43e-87 | 5 | 364 | 4 | 362 | Lipid A disaccharide synthetase [Cell wall/membrane/envelope biogenesis]. |
| pfam02684 | LpxB | 5.41e-66 | 5 | 351 | 1 | 346 | Lipid-A-disaccharide synthetase. This is a family of lipid-A-disaccharide synthetases, EC:2.4.2.128. These enzymes catalyze the reaction: UDP-2,3-bis(3-hydroxytetradecanoyl) glucosamine + 2,3-bis(3-hydroxytetradecanoyl)-beta-D-glucosaminyl 1-phosphate <=> UDP + 2,3-bis(3-hydroxytetradecanoyl)-D-glucosaminyl-1,6 -beta-D-2,3-bis(3-hydroxytetradecanoyl)-beta-D-glucosaminyl 1-phosphate. These enzymes catalyze the fist disaccharide step in the synthesis of lipid-A-disaccharide. |
| TIGR00215 | lpxB | 2.70e-59 | 2 | 347 | 5 | 348 | lipid-A-disaccharide synthase. Lipid-A precursor biosynthesis producing lipid A disaccharide in a condensation reaction. transcribed as part of an operon including lpxA [Cell envelope, Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides] |
| PRK14089 | PRK14089 | 7.57e-36 | 55 | 364 | 43 | 332 | lipid-A-disaccharide synthase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM45461.1 | 1.24e-212 | 1 | 383 | 10 | 392 |
| QSH40861.1 | 3.45e-163 | 2 | 385 | 5 | 388 |
| QHI67954.1 | 2.16e-63 | 1 | 377 | 1 | 369 |
| QUE51553.1 | 1.64e-59 | 1 | 328 | 1 | 321 |
| ABF41536.1 | 2.63e-59 | 5 | 385 | 4 | 380 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5W8N_A | 2.26e-28 | 5 | 378 | 9 | 376 | LipidA Disaccharide Synthase (LpxB)-6 solubilizing mutations [Escherichia coli BL21(DE3)] |
| 5W8S_A | 3.10e-28 | 5 | 378 | 9 | 376 | LipidA Disaccharide Synthase (LpxB)-7 solubilizing mutations [Escherichia coli BL21(DE3)],5W8X_A Lipid A Disaccharide Synthase (LpxB)-7 solubilizing mutations-Bound to UDP [Escherichia coli BL21(DE3)] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q1INL4 | 5.25e-60 | 5 | 385 | 4 | 380 | Lipid-A-disaccharide synthase OS=Koribacter versatilis (strain Ellin345) OX=204669 GN=lpxB PE=3 SV=1 |
| Q02AZ6 | 1.30e-51 | 5 | 375 | 4 | 366 | Lipid-A-disaccharide synthase OS=Solibacter usitatus (strain Ellin6076) OX=234267 GN=lpxB PE=3 SV=1 |
| C1F718 | 1.45e-51 | 4 | 373 | 6 | 381 | Lipid-A-disaccharide synthase OS=Acidobacterium capsulatum (strain ATCC 51196 / DSM 11244 / BCRC 80197 / JCM 7670 / NBRC 15755 / NCIMB 13165 / 161) OX=240015 GN=lpxB PE=3 SV=1 |
| Q2LVL8 | 2.15e-49 | 2 | 364 | 3 | 361 | Lipid-A-disaccharide synthase OS=Syntrophus aciditrophicus (strain SB) OX=56780 GN=lpxB PE=3 SV=1 |
| Q39T49 | 4.55e-48 | 2 | 370 | 6 | 369 | Lipid-A-disaccharide synthase OS=Geobacter metallireducens (strain ATCC 53774 / DSM 7210 / GS-15) OX=269799 GN=lpxB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
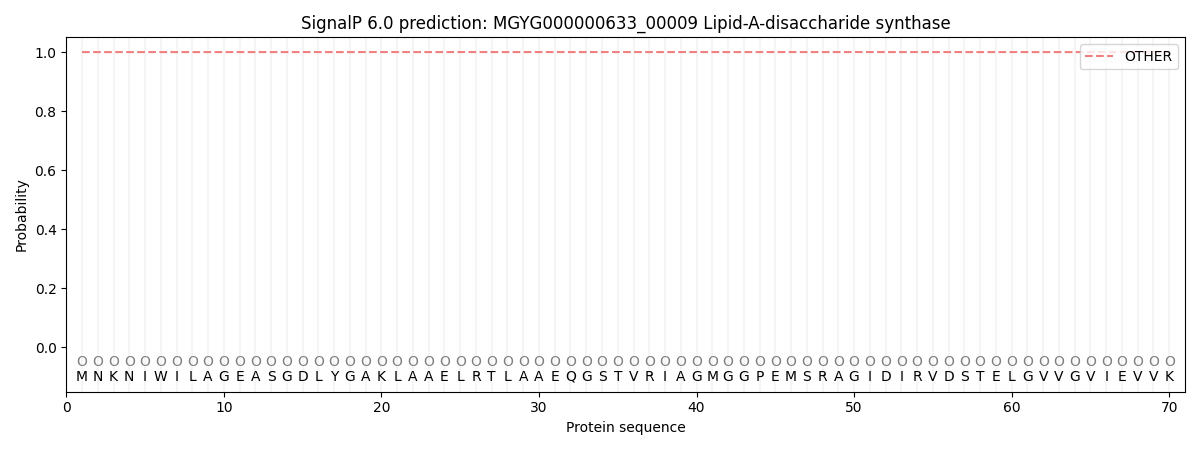
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000071 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
