You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000633_00758
You are here: Home > Sequence: MGYG000000633_00758
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Victivallis sp900768515 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; Victivallaceae; Victivallis; Victivallis sp900768515 | |||||||||||
| CAZyme ID | MGYG000000633_00758 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | Anti-sigma-I factor RsgI6 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 93156; End: 94490 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 64 | 382 | 7.4e-45 | 0.9570957095709571 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 8.06e-40 | 104 | 382 | 3 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 1.18e-36 | 82 | 380 | 22 | 306 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 2.99e-34 | 104 | 383 | 69 | 338 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM44415.1 | 1.28e-207 | 5 | 432 | 2 | 430 |
| AHF90707.1 | 2.81e-102 | 10 | 422 | 12 | 434 |
| QHV94149.1 | 1.44e-100 | 21 | 442 | 28 | 445 |
| QIP12711.1 | 1.81e-98 | 21 | 442 | 28 | 445 |
| QMW04637.1 | 2.09e-97 | 21 | 432 | 28 | 436 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1VBR_A | 5.44e-24 | 57 | 389 | 4 | 323 | Crystalstructure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBR_B Crystal structure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBU_A Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima],1VBU_B Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima] |
| 3NIY_A | 6.63e-24 | 57 | 389 | 20 | 339 | Crystalstructure of native xylanase 10B from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3NIY_B Crystal structure of native xylanase 10B from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3NJ3_A Crystal structure of xylanase 10B from Thermotoga petrophila RKU-1 in complex with xylobiose [Thermotoga petrophila RKU-1],3NJ3_B Crystal structure of xylanase 10B from Thermotoga petrophila RKU-1 in complex with xylobiose [Thermotoga petrophila RKU-1] |
| 6FHE_A | 1.23e-21 | 107 | 379 | 60 | 335 | Highlyactive enzymes by automated modular backbone assembly and sequence design [synthetic construct] |
| 7D88_A | 4.61e-20 | 42 | 427 | 62 | 407 | ChainA, Beta-xylanase [Bacillus sp. (in: Bacteria)] |
| 7D89_A | 8.35e-20 | 42 | 427 | 62 | 407 | ChainA, Beta-xylanase [Bacillus sp. (in: Bacteria)] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q60041 | 3.24e-18 | 104 | 389 | 72 | 342 | Endo-1,4-beta-xylanase B OS=Thermotoga neapolitana OX=2337 GN=xynB PE=3 SV=1 |
| P10478 | 2.61e-15 | 130 | 386 | 580 | 834 | Endo-1,4-beta-xylanase Z OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=xynZ PE=1 SV=3 |
| Q2PGV8 | 2.81e-14 | 130 | 387 | 95 | 356 | Endo-1,4-beta-xylanase 2 OS=Aureobasidium pullulans OX=5580 GN=xynII PE=1 SV=1 |
| P14768 | 4.72e-13 | 130 | 288 | 327 | 499 | Endo-1,4-beta-xylanase A OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=xynA PE=1 SV=2 |
| P07986 | 1.56e-12 | 130 | 379 | 105 | 346 | Exoglucanase/xylanase OS=Cellulomonas fimi OX=1708 GN=cex PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
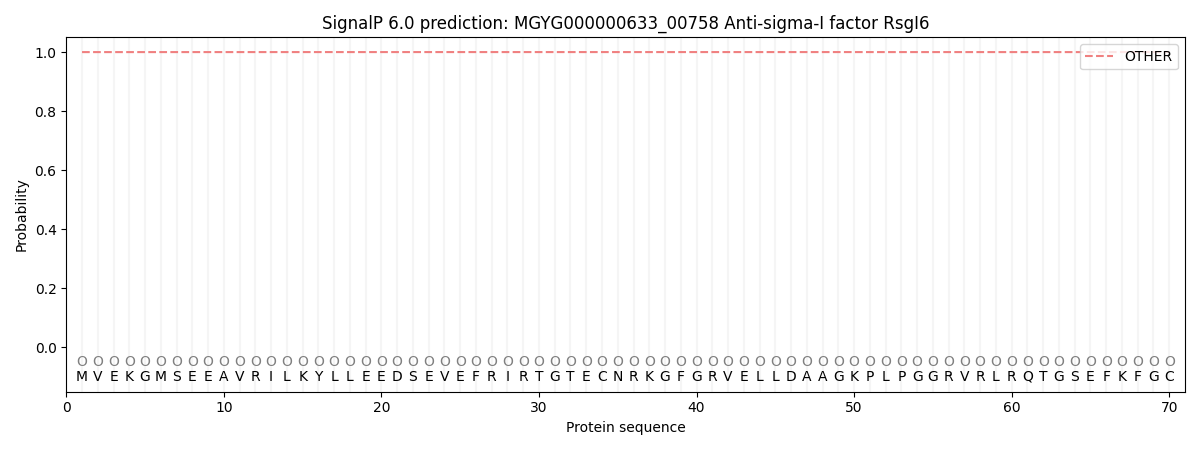
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000064 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
