You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000633_01137
Basic Information
help
| Species |
Victivallis sp900768515
|
| Lineage |
Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; Victivallaceae; Victivallis; Victivallis sp900768515
|
| CAZyme ID |
MGYG000000633_01137
|
| CAZy Family |
PL27 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
| Protein Length |
CGC |
Molecular Weight |
Isoelectric Point |
| 663 |
|
77327.99 |
6.2772 |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000000633 |
2845403 |
MAG |
Madagascar |
Africa |
|
| Gene Location |
Start: 46016;
End: 48007
Strand: -
|
No EC number prediction in MGYG000000633_01137.
| Family |
Start |
End |
Evalue |
family coverage |
| PL27 |
49 |
652 |
8.5e-150 |
0.9508196721311475 |
MGYG000000633_01137 has no CDD domain.
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
Description |
| 5NO8_A
|
2.76e-96 |
7 |
657 |
27 |
693 |
PolysaccharideLyase BACCELL_00875 [Bacteroides cellulosilyticus DSM 14838],5NO8_B Polysaccharide Lyase BACCELL_00875 [Bacteroides cellulosilyticus DSM 14838] |
| 5NOK_A
|
5.12e-90 |
7 |
657 |
27 |
693 |
PolysaccharideLyase BACCELL_00875 [Bacteroides cellulosilyticus DSM 14838],5NOK_B Polysaccharide Lyase BACCELL_00875 [Bacteroides cellulosilyticus DSM 14838] |
This protein is predicted as OTHER
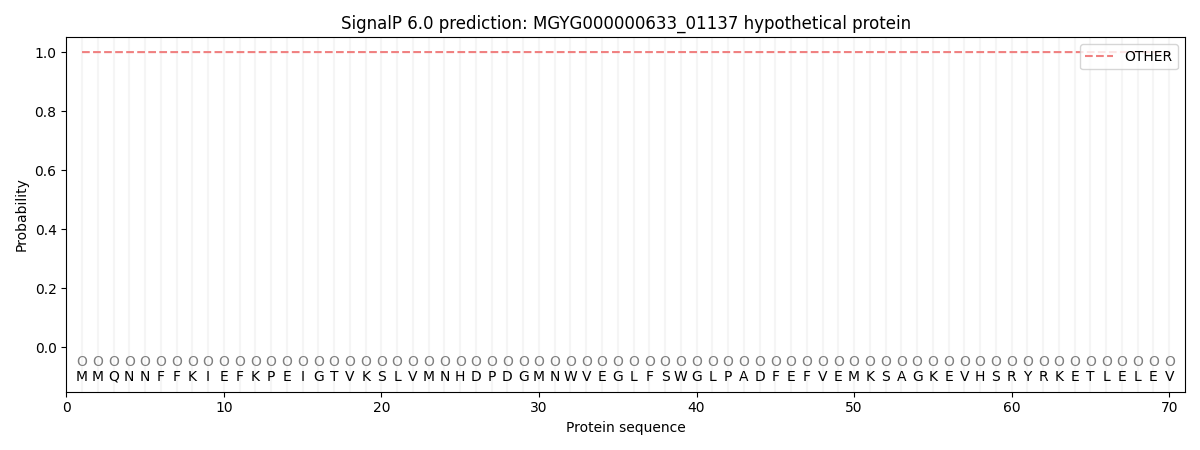
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000069
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000000633_01137.

