You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000644_00022
You are here: Home > Sequence: MGYG000000644_00022
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900546535 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900546535 | |||||||||||
| CAZyme ID | MGYG000000644_00022 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 35641; End: 38079 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 508 | 781 | 3.1e-43 | 0.6798679867986799 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 5.17e-35 | 508 | 779 | 54 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 1.59e-26 | 502 | 781 | 90 | 310 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 1.75e-25 | 508 | 777 | 120 | 335 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
| pfam02018 | CBM_4_9 | 1.62e-12 | 329 | 446 | 8 | 123 | Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. |
| pfam00331 | Glyco_hydro_10 | 2.74e-08 | 55 | 141 | 1 | 85 | Glycosyl hydrolase family 10. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT92890.1 | 1.43e-212 | 1 | 802 | 1 | 763 |
| ALJ61540.1 | 1.14e-211 | 1 | 802 | 1 | 763 |
| EDV05054.1 | 7.47e-183 | 1 | 812 | 1 | 782 |
| QDO69424.1 | 4.19e-182 | 1 | 812 | 1 | 782 |
| QCP72441.1 | 1.26e-171 | 1 | 803 | 1 | 789 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6LPS_A | 3.02e-16 | 600 | 776 | 170 | 346 | ChainA, Beta-xylanase [Halalkalibacterium halodurans] |
| 7CPL_A | 3.32e-16 | 600 | 776 | 171 | 347 | XylanaseR from Bacillus sp. TAR-1 [Bacillus sp. TAR1] |
| 7CPK_A | 3.32e-16 | 600 | 776 | 171 | 347 | XylanaseR from Bacillus sp. TAR-1 [Bacillus sp. TAR1] |
| 2UWF_A | 1.02e-15 | 600 | 776 | 169 | 345 | ChainA, ALKALINE ACTIVE ENDOXYLANASE [Halalkalibacterium halodurans] |
| 3W24_A | 1.55e-14 | 600 | 788 | 166 | 333 | Thehigh-resolution crystal structure of TsXylA, intracellular xylanase from Thermoanaerobacterium saccharolyticum JW/SL-YS485 [Thermoanaerobacterium saccharolyticum JW/SL-YS485] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P07528 | 2.53e-15 | 600 | 776 | 215 | 391 | Endo-1,4-beta-xylanase A OS=Alkalihalobacillus halodurans (strain ATCC BAA-125 / DSM 18197 / FERM 7344 / JCM 9153 / C-125) OX=272558 GN=xynA PE=1 SV=1 |
| P38535 | 1.78e-12 | 600 | 781 | 367 | 526 | Exoglucanase XynX OS=Acetivibrio thermocellus OX=1515 GN=xynX PE=3 SV=1 |
| P36917 | 2.40e-12 | 600 | 781 | 515 | 674 | Endo-1,4-beta-xylanase A OS=Thermoanaerobacterium saccharolyticum OX=28896 GN=xynA PE=1 SV=1 |
| Q2PGV8 | 3.47e-11 | 605 | 776 | 185 | 348 | Endo-1,4-beta-xylanase 2 OS=Aureobasidium pullulans OX=5580 GN=xynII PE=1 SV=1 |
| P40944 | 8.52e-11 | 600 | 781 | 512 | 677 | Endo-1,4-beta-xylanase A OS=Caldicellulosiruptor sp. (strain Rt8B.4) OX=28238 GN=xynA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
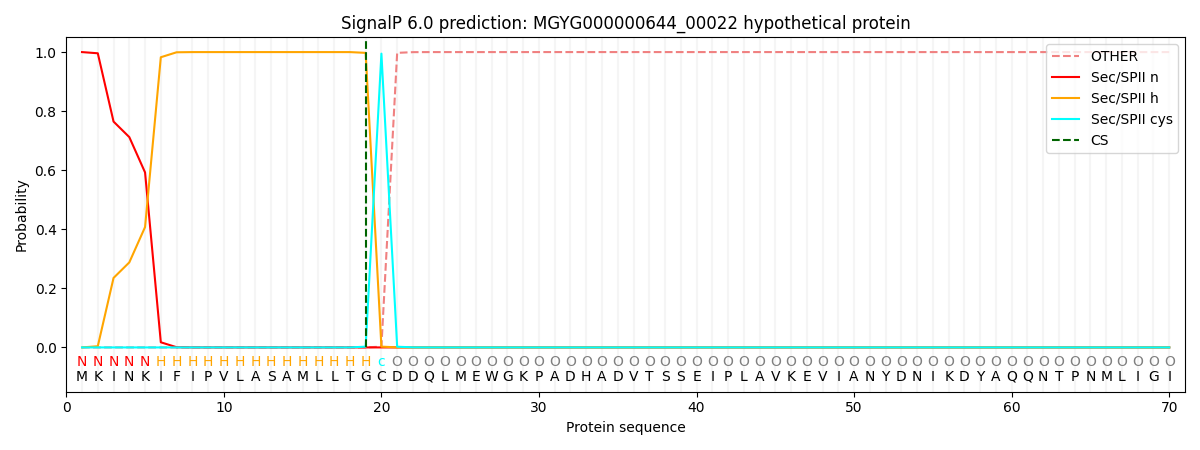
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000065 | 0.000000 | 0.000000 | 0.000000 |
