You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000644_00501
You are here: Home > Sequence: MGYG000000644_00501
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900546535 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900546535 | |||||||||||
| CAZyme ID | MGYG000000644_00501 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 68630; End: 70504 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 299 | 467 | 1.9e-41 | 0.8217821782178217 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3866 | PelB | 2.09e-50 | 159 | 581 | 1 | 345 | Pectate lyase [Carbohydrate transport and metabolism]. |
| smart00656 | Amb_all | 4.45e-28 | 302 | 468 | 17 | 190 | Amb_all domain. |
| pfam00544 | Pec_lyase_C | 1.94e-14 | 308 | 464 | 42 | 211 | Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNT67442.1 | 0.0 | 1 | 622 | 1 | 614 |
| QCD40816.1 | 2.06e-211 | 40 | 589 | 1 | 558 |
| QCP73706.1 | 2.06e-211 | 40 | 589 | 1 | 558 |
| QUT73893.1 | 2.97e-202 | 105 | 622 | 20 | 543 |
| ATA84997.1 | 3.66e-198 | 106 | 613 | 39 | 541 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3VMV_A | 5.53e-25 | 252 | 538 | 33 | 317 | Crystalstructure of pectate lyase Bsp165PelA from Bacillus sp. N165 [Bacillus sp. N16-5],3VMW_A Crystal structure of pectate lyase Bsp165PelA from Bacillus sp. N165 in complex with trigalacturonate [Bacillus sp. N16-5] |
| 1AIR_A | 1.87e-18 | 225 | 579 | 15 | 342 | ChainA, PECTATE LYASE C [Dickeya chrysanthemi],1O88_A Chain A, Pectate Lyase C [Dickeya chrysanthemi],1O8D_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8E_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8F_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8G_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8H_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8I_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8J_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8K_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8L_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8M_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1PLU_A Chain A, Protein (pectate Lyase C) [Dickeya chrysanthemi],2PEC_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi] |
| 2EWE_A | 4.53e-18 | 225 | 579 | 15 | 342 | ChainA, Pectate lyase C [Dickeya chrysanthemi] |
| 1VBL_A | 1.46e-17 | 205 | 542 | 43 | 411 | Structureof the thermostable pectate lyase PL 47 [Bacillus sp. TS-47] |
| 5AMV_A | 6.69e-10 | 302 | 456 | 128 | 303 | Structuralinsights into the loss of catalytic competence in pectate lyase at low pH [Bacillus subtilis],5X2I_A Polygalacturonate Lyase by Fusing with a Self-assembling Amphipathic Peptide [Bacillus subtilis subsp. subtilis str. 168] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B1B6T1 | 1.18e-21 | 200 | 473 | 40 | 281 | Pectate trisaccharide-lyase OS=Bacillus sp. OX=1409 GN=pel PE=1 SV=1 |
| Q65DC2 | 1.18e-21 | 200 | 473 | 40 | 281 | Pectate trisaccharide-lyase OS=Bacillus licheniformis (strain ATCC 14580 / DSM 13 / JCM 2505 / CCUG 7422 / NBRC 12200 / NCIMB 9375 / NCTC 10341 / NRRL NRS-1264 / Gibson 46) OX=279010 GN=BLi04129 PE=3 SV=1 |
| Q8GCB2 | 1.18e-21 | 200 | 473 | 40 | 281 | Pectate trisaccharide-lyase OS=Bacillus licheniformis OX=1402 GN=pelA PE=1 SV=1 |
| P11073 | 1.30e-17 | 225 | 579 | 37 | 364 | Pectate lyase C OS=Dickeya chrysanthemi OX=556 GN=pelC PE=1 SV=1 |
| Q5AVN4 | 1.53e-15 | 302 | 481 | 99 | 281 | Pectate lyase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
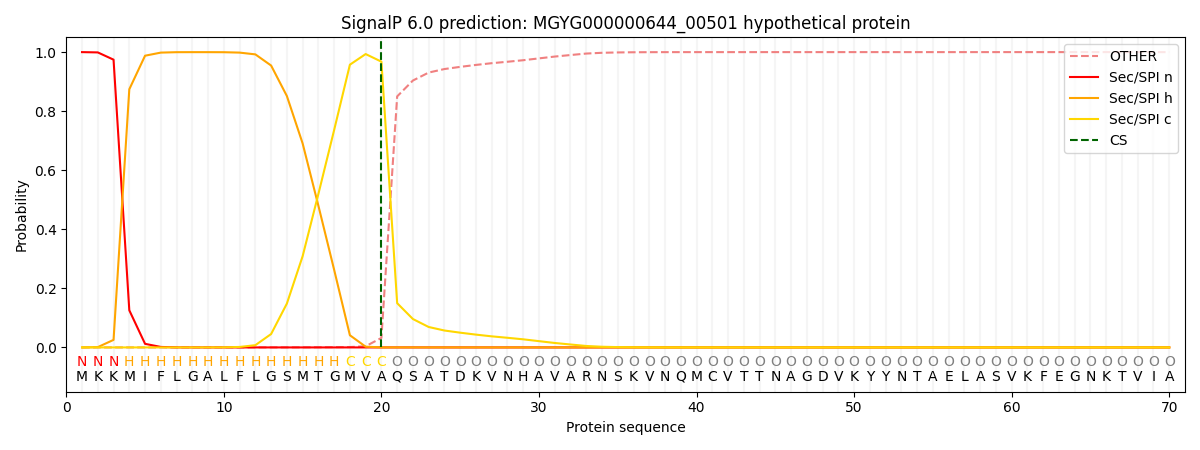
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000737 | 0.998385 | 0.000282 | 0.000207 | 0.000181 | 0.000165 |
