You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000658_00883
You are here: Home > Sequence: MGYG000000658_00883
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotellamassilia; | |||||||||||
| CAZyme ID | MGYG000000658_00883 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | Endo-1,4-beta-xylanase Z | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 30144; End: 31025 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 38 | 285 | 1.2e-53 | 0.986784140969163 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00756 | Esterase | 3.35e-33 | 38 | 282 | 1 | 245 | Putative esterase. This family contains Esterase D. However it is not clear if all members of the family have the same function. This family is related to the pfam00135 family. |
| COG2382 | Fes | 9.37e-27 | 38 | 286 | 76 | 298 | Enterochelin esterase or related enzyme [Inorganic ion transport and metabolism]. |
| COG0627 | FrmB | 5.08e-24 | 60 | 287 | 51 | 312 | S-formylglutathione hydrolase FrmB [Defense mechanisms]. |
| COG2819 | YbbA | 7.10e-08 | 30 | 190 | 8 | 176 | Predicted hydrolase of the alpha/beta superfamily [General function prediction only]. |
| COG4947 | COG4947 | 3.29e-04 | 120 | 276 | 62 | 223 | Esterase/lipase superfamily enzyme [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM58192.1 | 5.80e-26 | 45 | 272 | 162 | 385 |
| QJW99051.1 | 6.14e-26 | 27 | 285 | 269 | 507 |
| ADV60895.1 | 6.97e-26 | 38 | 283 | 144 | 384 |
| AYB44329.1 | 8.02e-25 | 23 | 277 | 13 | 249 |
| QQA30430.1 | 1.05e-24 | 45 | 272 | 162 | 385 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5VOL_A | 1.57e-99 | 24 | 287 | 5 | 272 | Bacint_04212ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_B Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_C Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_D Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_E Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_F Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_G Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_H Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393] |
| 6RZN_A | 1.86e-22 | 37 | 283 | 138 | 384 | Crystalstructure of the N-terminal carbohydrate binding module family 48 and ferulic acid esterase from the multi-enzyme CE1-GH62-GH10 [uncultured bacterium],6RZN_B Crystal structure of the N-terminal carbohydrate binding module family 48 and ferulic acid esterase from the multi-enzyme CE1-GH62-GH10 [uncultured bacterium] |
| 7B5V_A | 1.43e-17 | 38 | 162 | 146 | 275 | ChainA, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B5V_B Chain B, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B6B_A Chain A, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B6B_B Chain B, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836] |
| 1JJF_A | 2.07e-17 | 18 | 286 | 10 | 260 | ChainA, Endo-1,4-beta-xylanase Z [Acetivibrio thermocellus] |
| 6RZO_A | 4.92e-17 | 43 | 176 | 129 | 260 | Crystalstructure of the N-terminal carbohydrate binding module family 48 and ferulic acid esterase from the multi-enzyme CE1-GH62-GH10 [uncultured bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P10478 | 2.12e-16 | 7 | 286 | 18 | 279 | Endo-1,4-beta-xylanase Z OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=xynZ PE=1 SV=3 |
| P0C1D6 | 1.06e-12 | 28 | 286 | 102 | 385 | Protein PS1 OS=Corynebacterium glutamicum (strain ATCC 13032 / DSM 20300 / BCRC 11384 / JCM 1318 / LMG 3730 / NCIMB 10025) OX=196627 GN=csp1 PE=1 SV=1 |
| D5EY13 | 2.68e-12 | 35 | 176 | 504 | 646 | Endo-1,4-beta-xylanase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=xyn10D-fae1A PE=1 SV=1 |
| D5EXZ4 | 6.30e-12 | 43 | 283 | 443 | 668 | Carbohydrate acetyl esterase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=axe1-6A PE=1 SV=1 |
| P9WM38 | 1.37e-09 | 48 | 267 | 191 | 425 | Esterase MT1326 OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=MT1326 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
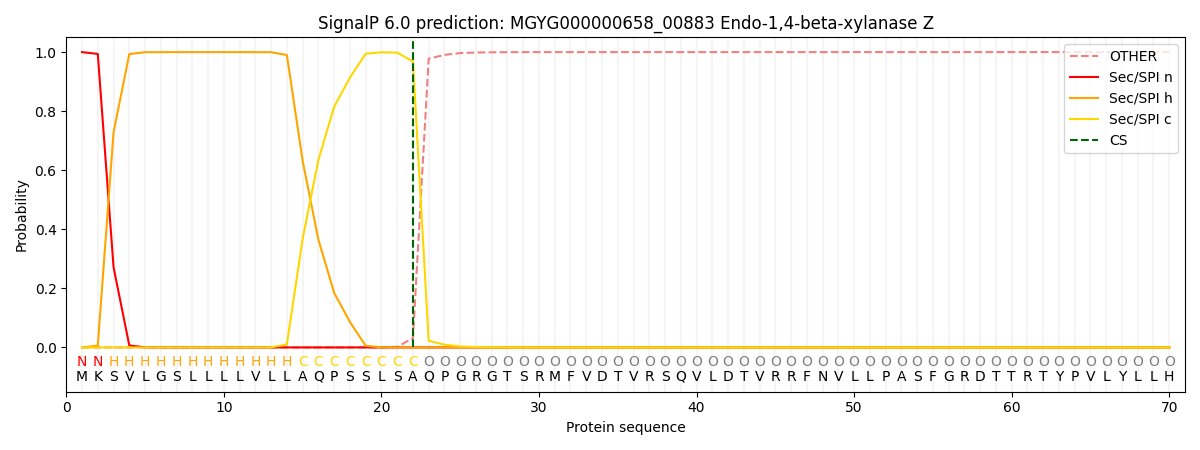
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000205 | 0.999163 | 0.000160 | 0.000160 | 0.000146 | 0.000141 |
