You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000659_01128
You are here: Home > Sequence: MGYG000000659_01128
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA10281 sp900556195 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; Borkfalkiaceae; UBA10281; UBA10281 sp900556195 | |||||||||||
| CAZyme ID | MGYG000000659_01128 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4565; End: 7438 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 131 | 797 | 3.6e-67 | 0.648936170212766 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 2.04e-31 | 142 | 491 | 66 | 399 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10340 | ebgA | 1.77e-16 | 135 | 459 | 105 | 400 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| PRK10150 | PRK10150 | 6.80e-11 | 161 | 566 | 88 | 482 | beta-D-glucuronidase; Provisional |
| PRK09525 | lacZ | 1.15e-08 | 338 | 453 | 296 | 408 | beta-galactosidase. |
| pfam00703 | Glyco_hydro_2 | 4.15e-07 | 266 | 374 | 2 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AEE97824.1 | 2.32e-232 | 34 | 939 | 21 | 1043 |
| QBE96176.1 | 4.44e-228 | 51 | 777 | 24 | 814 |
| QUI23767.1 | 2.77e-226 | 33 | 789 | 4 | 817 |
| QNK56944.1 | 1.17e-218 | 92 | 806 | 103 | 852 |
| QDH23079.1 | 3.00e-217 | 80 | 812 | 59 | 839 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6MVF_A | 7.90e-18 | 168 | 594 | 100 | 492 | Crystalstructure of FMN-binding beta-glucuronidase from Facaelibacterium prausnitzii L2-6 [Faecalibacterium prausnitzii L2-6],6MVF_B Crystal structure of FMN-binding beta-glucuronidase from Facaelibacterium prausnitzii L2-6 [Faecalibacterium prausnitzii L2-6],6MVF_C Crystal structure of FMN-binding beta-glucuronidase from Facaelibacterium prausnitzii L2-6 [Faecalibacterium prausnitzii L2-6],6MVF_D Crystal structure of FMN-binding beta-glucuronidase from Facaelibacterium prausnitzii L2-6 [Faecalibacterium prausnitzii L2-6],6MVF_E Crystal structure of FMN-binding beta-glucuronidase from Facaelibacterium prausnitzii L2-6 [Faecalibacterium prausnitzii L2-6],6MVF_F Crystal structure of FMN-binding beta-glucuronidase from Facaelibacterium prausnitzii L2-6 [Faecalibacterium prausnitzii L2-6] |
| 3OB8_A | 1.76e-15 | 135 | 464 | 108 | 425 | Structureof the beta-galactosidase from Kluyveromyces lactis in complex with galactose [Kluyveromyces lactis],3OB8_B Structure of the beta-galactosidase from Kluyveromyces lactis in complex with galactose [Kluyveromyces lactis],3OB8_C Structure of the beta-galactosidase from Kluyveromyces lactis in complex with galactose [Kluyveromyces lactis],3OB8_D Structure of the beta-galactosidase from Kluyveromyces lactis in complex with galactose [Kluyveromyces lactis],3OBA_A Structure of the beta-galactosidase from Kluyveromyces lactis [Kluyveromyces lactis],3OBA_B Structure of the beta-galactosidase from Kluyveromyces lactis [Kluyveromyces lactis],3OBA_C Structure of the beta-galactosidase from Kluyveromyces lactis [Kluyveromyces lactis],3OBA_D Structure of the beta-galactosidase from Kluyveromyces lactis [Kluyveromyces lactis] |
| 7KGY_A | 7.94e-15 | 155 | 508 | 86 | 408 | ChainA, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_B Chain B, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_C Chain C, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_D Chain D, Beta-glucuronidase [Faecalibacterium prausnitzii] |
| 7VQM_A | 1.04e-14 | 143 | 594 | 98 | 503 | ChainA, GH2 beta-galacturonate AqGalA [Aquimarina sp.],7VQM_B Chain B, GH2 beta-galacturonate AqGalA [Aquimarina sp.],7VQM_C Chain C, GH2 beta-galacturonate AqGalA [Aquimarina sp.],7VQM_D Chain D, GH2 beta-galacturonate AqGalA [Aquimarina sp.] |
| 6SD0_A | 6.24e-14 | 129 | 455 | 104 | 387 | Structureof beta-galactosidase from Thermotoga maritima. [Thermotoga maritima MSB8],6SD0_B Structure of beta-galactosidase from Thermotoga maritima. [Thermotoga maritima MSB8],6SD0_C Structure of beta-galactosidase from Thermotoga maritima. [Thermotoga maritima MSB8],6SD0_D Structure of beta-galactosidase from Thermotoga maritima. [Thermotoga maritima MSB8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KPJ7 | 5.31e-21 | 152 | 701 | 117 | 626 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
| O52847 | 1.95e-20 | 130 | 511 | 124 | 491 | Beta-galactosidase OS=Priestia megaterium (strain DSM 319 / IMG 1521) OX=592022 GN=bgaM PE=3 SV=1 |
| P24131 | 6.48e-17 | 130 | 505 | 112 | 465 | Beta-galactosidase OS=Clostridium acetobutylicum OX=1488 GN=cbgA PE=2 SV=2 |
| P77989 | 2.87e-16 | 140 | 489 | 57 | 378 | Beta-galactosidase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=lacZ PE=3 SV=2 |
| P23989 | 6.28e-16 | 124 | 505 | 105 | 464 | Beta-galactosidase OS=Streptococcus thermophilus OX=1308 GN=lacZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
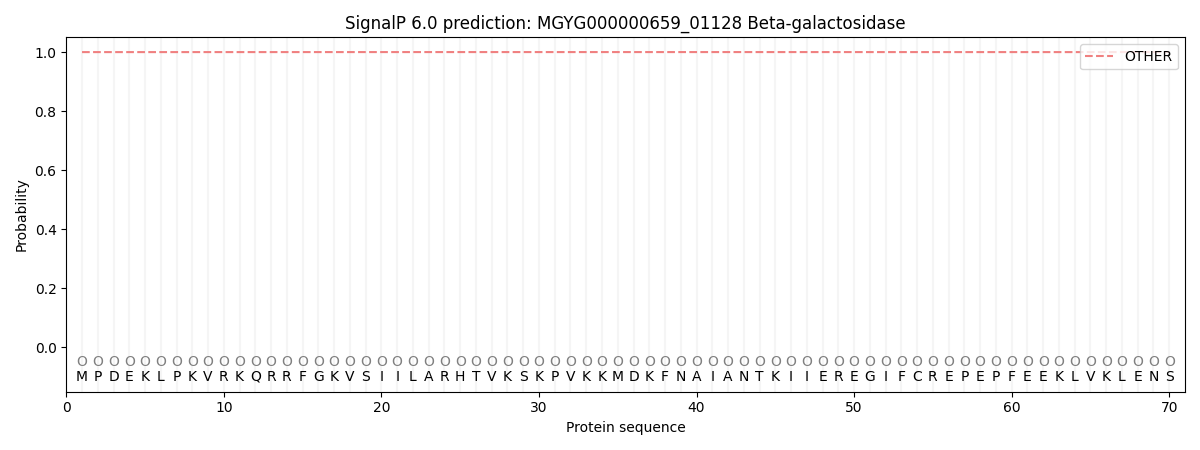
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000070 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
