You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000666_00333
You are here: Home > Sequence: MGYG000000666_00333
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; UBA1829; ; | |||||||||||
| CAZyme ID | MGYG000000666_00333 | |||||||||||
| CAZy Family | GT5 | |||||||||||
| CAZyme Description | Glycogen synthase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 32290; End: 33039 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam08323 | Glyco_transf_5 | 2.42e-105 | 3 | 243 | 1 | 238 | Starch synthase catalytic domain. |
| PRK00654 | glgA | 4.77e-101 | 1 | 245 | 1 | 228 | glycogen synthase GlgA. |
| cd03791 | GT5_Glycogen_synthase_DULL1-like | 1.29e-99 | 2 | 245 | 1 | 239 | Glycogen synthase GlgA and similar proteins. This family is most closely related to the GT5 family of glycosyltransferases. Glycogen synthase (EC:2.4.1.21) catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. |
| COG0297 | GlgA | 1.59e-75 | 1 | 249 | 1 | 244 | Glycogen synthase [Carbohydrate transport and metabolism]. |
| PRK14099 | PRK14099 | 2.34e-45 | 1 | 245 | 4 | 240 | glycogen synthase GlgA. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ABM04087.1 | 9.80e-110 | 1 | 247 | 1 | 248 |
| QDU62030.1 | 2.17e-68 | 1 | 247 | 1 | 246 |
| QDV52381.1 | 2.53e-63 | 1 | 242 | 1 | 245 |
| QDT44223.1 | 6.89e-63 | 1 | 242 | 1 | 245 |
| QEG18479.1 | 2.69e-62 | 1 | 244 | 1 | 247 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1RZU_A | 8.98e-38 | 1 | 248 | 1 | 240 | ChainA, Glycogen synthase 1 [Agrobacterium tumefaciens],1RZU_B Chain B, Glycogen synthase 1 [Agrobacterium tumefaciens] |
| 3D1J_A | 3.01e-37 | 1 | 245 | 1 | 235 | ChainA, Glycogen synthase [Escherichia coli] |
| 2QZS_A | 6.44e-37 | 1 | 245 | 1 | 235 | CrystalStructure of Wild-type E.coli GS in complex with ADP and Glucose(wtGSb) [Escherichia coli],2R4T_A Crystal Structure of Wild-type E.coli GS in Complex with ADP and Glucose(wtGSc) [Escherichia coli],2R4U_A Crystal Structure of Wild-type E.coli GS in complex with ADP and Glucose(wtGSd) [Escherichia coli],3GUH_A Crystal Structure of Wild-type E.coli GS in complex with ADP and DGM [Escherichia coli K-12] |
| 3COP_A | 6.44e-37 | 1 | 245 | 1 | 235 | ChainA, Glycogen synthase [Escherichia coli],3CX4_A Chain A, Glycogen synthase [Escherichia coli] |
| 1RZV_A | 8.86e-36 | 2 | 248 | 2 | 240 | ChainA, Glycogen synthase 1 [Agrobacterium tumefaciens],1RZV_B Chain B, Glycogen synthase 1 [Agrobacterium tumefaciens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q74ED9 | 7.04e-59 | 1 | 247 | 4 | 250 | Glycogen synthase 1 OS=Geobacter sulfurreducens (strain ATCC 51573 / DSM 12127 / PCA) OX=243231 GN=glgA1 PE=3 SV=1 |
| B5YG91 | 4.96e-58 | 1 | 245 | 1 | 256 | Glycogen synthase OS=Thermodesulfovibrio yellowstonii (strain ATCC 51303 / DSM 11347 / YP87) OX=289376 GN=glgA PE=3 SV=1 |
| Q7UPY2 | 7.88e-54 | 1 | 243 | 1 | 251 | Glycogen synthase OS=Rhodopirellula baltica (strain DSM 10527 / NCIMB 13988 / SH1) OX=243090 GN=glgA PE=3 SV=1 |
| Q1DCS0 | 1.21e-53 | 1 | 245 | 1 | 243 | Glycogen synthase OS=Myxococcus xanthus (strain DK1622) OX=246197 GN=glgA PE=3 SV=1 |
| A7H6L1 | 2.28e-53 | 1 | 245 | 1 | 237 | Glycogen synthase OS=Anaeromyxobacter sp. (strain Fw109-5) OX=404589 GN=glgA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
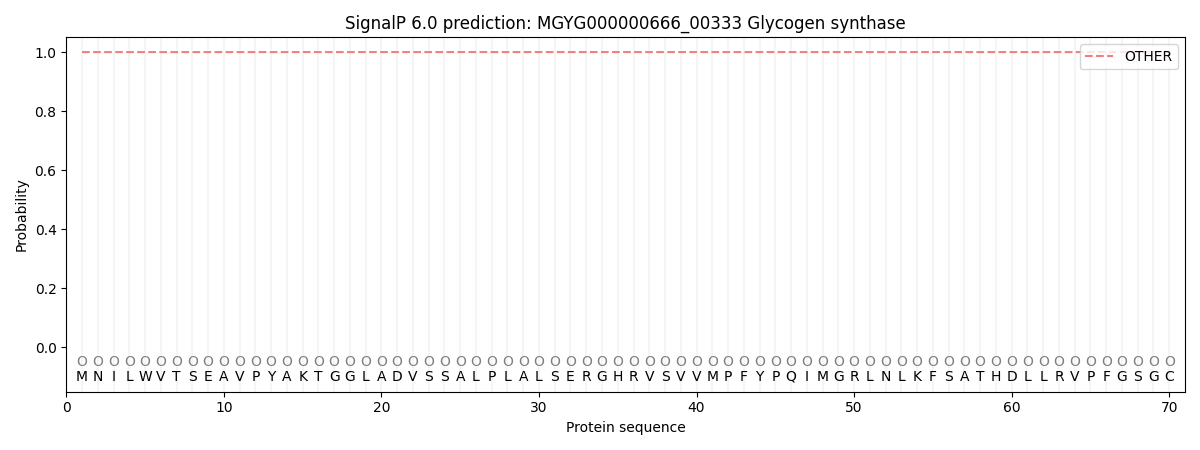
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000060 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
