You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000670_00155
You are here: Home > Sequence: MGYG000000670_00155
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Faecalibacillus sp900544435 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Erysipelotrichales; Erysipelatoclostridiaceae; Faecalibacillus; Faecalibacillus sp900544435 | |||||||||||
| CAZyme ID | MGYG000000670_00155 | |||||||||||
| CAZy Family | GH170 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 144701; End: 145870 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH170 | 5 | 252 | 1.4e-71 | 0.6885714285714286 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam19200 | DUF871_N | 2.15e-82 | 6 | 239 | 1 | 233 | DUF871 N-terminal domain. This family consists of several conserved hypothetical proteins from bacteria and archaea. The function of this family is unknown. |
| COG3589 | COG3589 | 5.76e-79 | 3 | 389 | 1 | 358 | Uncharacterized protein [Function unknown]. |
| pfam05913 | DUF871 | 1.52e-29 | 271 | 388 | 1 | 116 | Bacterial protein of unknown function (DUF871). This family consists of several conserved hypothetical proteins from bacteria and archaea. The function of this family is unknown. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCL57950.1 | 4.50e-170 | 99 | 332 | 1 | 234 |
| AMK55490.1 | 5.69e-170 | 1 | 389 | 15 | 408 |
| QJA01274.1 | 9.16e-103 | 6 | 389 | 4 | 364 |
| QAT39807.1 | 3.79e-102 | 6 | 389 | 4 | 362 |
| QIX07901.1 | 2.37e-100 | 6 | 389 | 4 | 364 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1X7F_A | 8.75e-75 | 6 | 389 | 29 | 385 | Crystalstructure of an uncharacterized B. cereus protein [Bacillus cereus ATCC 14579] |
| 2P0O_A | 2.13e-15 | 7 | 388 | 6 | 358 | Crystalstructure of a conserved protein from locus EF_2437 in Enterococcus faecalis with an unknown function [Enterococcus faecalis V583] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
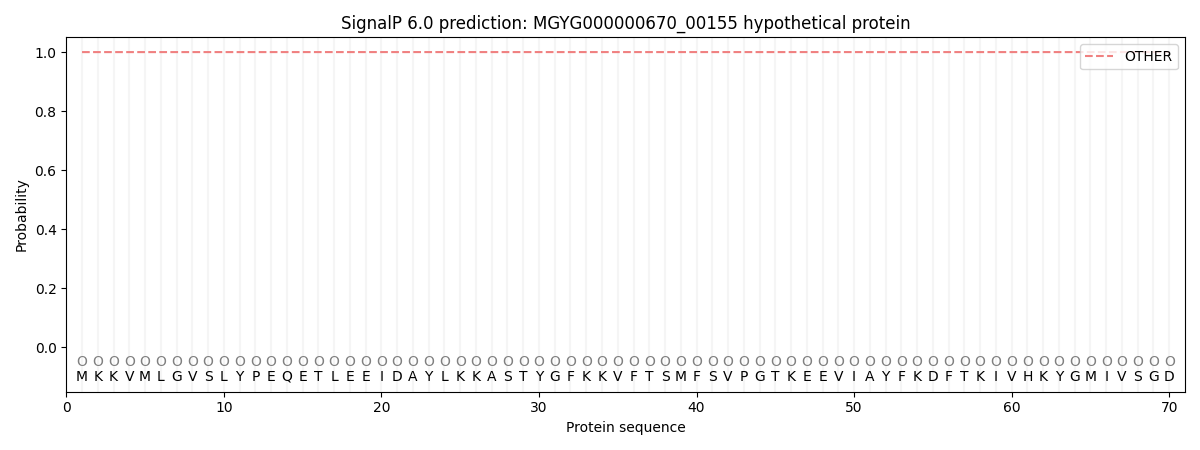
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000056 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
