You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000675_01014
You are here: Home > Sequence: MGYG000000675_01014
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides congonensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides congonensis | |||||||||||
| CAZyme ID | MGYG000000675_01014 | |||||||||||
| CAZy Family | GT11 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 66268; End: 68415 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT11 | 15 | 264 | 3.7e-55 | 0.9601449275362319 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd11301 | Fut1_Fut2_like | 3.85e-41 | 1 | 244 | 1 | 254 | Alpha-1,2-fucosyltransferase. Alpha-1,2-fucosyltransferases (Fut1, Fut2) catalyze the transfer of alpha-L-fucose to the terminal beta-D-galactose residue of glycoconjugates via an alpha-1,2-linkage, generating carbohydrate structures that exhibit H-antigenicity for blood-group carbohydrates. These structures also act as ligands for morphogenesis, the adhesion of microbes, and metastasizing cancer cells. Fut1 is responsible for producing the H antigen on red blood cells. Fut2 is expressed in epithelia of secretory tissues, and individuals termed "secretors" have at least one functional copy of the gene; they secrete H antigen which is further processed into A and/or B antigens depending on the ABO genotype. O-fucosyltransferase-like proteins are GDP-fucose dependent enzymes with similarities to the family 1 glycosyltransferases (GT1). They are soluble ER proteins that may be proteolytically cleaved from a membrane-associated preprotein, and are involved in the O-fucosylation of protein substrates, the core fucosylation of growth factor receptors, and other processes. |
| pfam01531 | Glyco_transf_11 | 1.66e-22 | 16 | 264 | 41 | 295 | Glycosyl transferase family 11. This family contains several fucosyl transferase enzymes. |
| COG3306 | COG3306 | 8.47e-07 | 501 | 575 | 2 | 100 | Glycosyltransferase involved in LPS biosynthesis, GR25 family [Cell wall/membrane/envelope biogenesis]. |
| cd06532 | Glyco_transf_25 | 1.07e-06 | 503 | 571 | 1 | 92 | Glycosyltransferase family 25 [lipooligosaccharide (LOS) biosynthesis protein] is a family of glycosyltransferases involved in LOS biosynthesis. The members include the beta(1,4) galactosyltransferases: Lgt2 of Moraxella catarrhalis, LgtB and LgtE of Neisseria gonorrhoeae and Lic2A of Haemophilus influenzae. M. catarrhalis Lgt2 catalyzes the addition of galactose (Gal) to the growing chain of LOS on the cell surface. N. gonorrhoeae LgtB and LgtE link Gal-beta(1,4) to GlcNAc (N-acetylglucosamine) and Glc (glucose), respectively. The genes encoding LgtB and LgtE are two genes of a five gene locus involved in the synthesis of gonococcal LOS. LgtE is believed to perform the first step in LOS biosynthesis. |
| cd06532 | Glyco_transf_25 | 0.002 | 276 | 344 | 1 | 92 | Glycosyltransferase family 25 [lipooligosaccharide (LOS) biosynthesis protein] is a family of glycosyltransferases involved in LOS biosynthesis. The members include the beta(1,4) galactosyltransferases: Lgt2 of Moraxella catarrhalis, LgtB and LgtE of Neisseria gonorrhoeae and Lic2A of Haemophilus influenzae. M. catarrhalis Lgt2 catalyzes the addition of galactose (Gal) to the growing chain of LOS on the cell surface. N. gonorrhoeae LgtB and LgtE link Gal-beta(1,4) to GlcNAc (N-acetylglucosamine) and Glc (glucose), respectively. The genes encoding LgtB and LgtE are two genes of a five gene locus involved in the synthesis of gonococcal LOS. LgtE is believed to perform the first step in LOS biosynthesis. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CDN30683.1 | 1.07e-80 | 92 | 705 | 364 | 991 |
| QUB60703.1 | 7.79e-77 | 495 | 711 | 370 | 586 |
| QUB69923.1 | 7.94e-76 | 495 | 711 | 370 | 586 |
| ATR96316.1 | 1.43e-75 | 502 | 710 | 382 | 590 |
| QUB82181.1 | 4.17e-75 | 495 | 711 | 370 | 586 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q58YV9 | 5.57e-16 | 94 | 264 | 124 | 300 | O-antigen biosynthesis glycosyltransferase WbnK OS=Escherichia coli OX=562 GN=wbnK PE=1 SV=1 |
| P91200 | 8.58e-09 | 16 | 263 | 74 | 341 | Galactoside 2-alpha-L-fucosyltransferase OS=Caenorhabditis elegans OX=6239 GN=fut-2 PE=1 SV=1 |
| Q966A0 | 1.48e-06 | 16 | 263 | 76 | 348 | Putative galactoside 2-alpha-L-fucosyltransferase svh-11 OS=Caenorhabditis elegans OX=6239 GN=svh-11 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
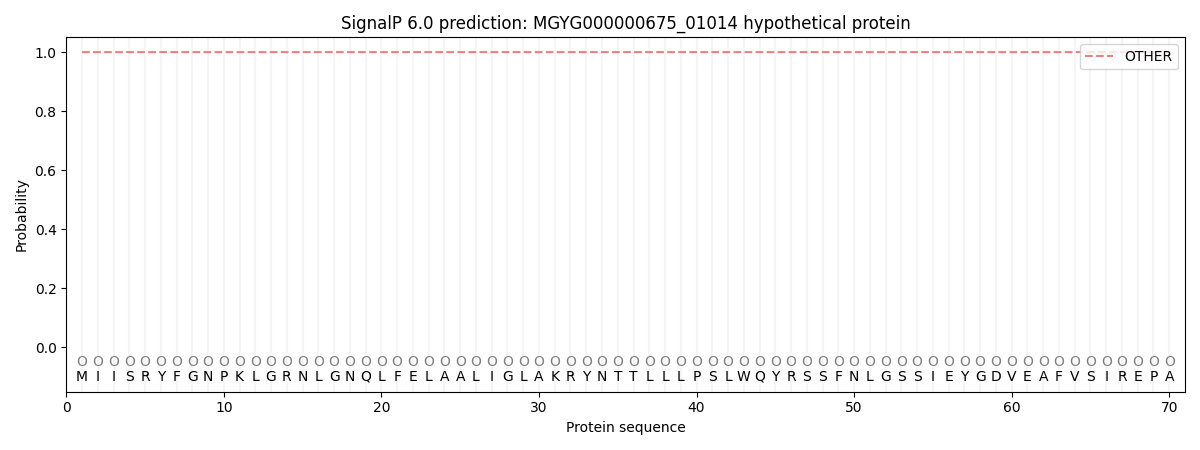
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000015 | 0.000016 | 0.000001 | 0.000000 | 0.000000 | 0.000000 |
