You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000678_00512
You are here: Home > Sequence: MGYG000000678_00512
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-485 sp002491165 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; CAG-485; CAG-485 sp002491165 | |||||||||||
| CAZyme ID | MGYG000000678_00512 | |||||||||||
| CAZy Family | GH35 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7781; End: 10162 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH35 | 46 | 357 | 1.1e-116 | 0.990228013029316 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01301 | Glyco_hydro_35 | 1.67e-155 | 45 | 355 | 1 | 313 | Glycosyl hydrolases family 35. |
| PLN03059 | PLN03059 | 5.71e-53 | 10 | 620 | 9 | 715 | beta-galactosidase; Provisional |
| COG1874 | GanA | 6.50e-42 | 44 | 621 | 6 | 593 | Beta-galactosidase GanA [Carbohydrate transport and metabolism]. |
| pfam02449 | Glyco_hydro_42 | 1.10e-12 | 62 | 230 | 3 | 176 | Beta-galactosidase. This group of beta-galactosidase enzymes belong to the glycosyl hydrolase 42 family. The enzyme catalyzes the hydrolysis of terminal, non-reducing terminal beta-D-galactosidase residues. |
| pfam00754 | F5_F8_type_C | 4.95e-05 | 724 | 790 | 5 | 69 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCD35705.1 | 0.0 | 38 | 789 | 346 | 1096 |
| QUT89998.1 | 0.0 | 32 | 789 | 343 | 1097 |
| ALJ58885.1 | 0.0 | 32 | 789 | 343 | 1097 |
| QDO67465.1 | 0.0 | 32 | 789 | 343 | 1097 |
| QUU00371.1 | 0.0 | 35 | 789 | 347 | 1097 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6EON_A | 1.79e-310 | 10 | 791 | 7 | 772 | GalactanaseBT0290 [Bacteroides thetaiotaomicron VPI-5482] |
| 3D3A_A | 3.84e-257 | 33 | 642 | 2 | 596 | Crystalstructure of a beta-galactosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482] |
| 4MAD_A | 7.11e-146 | 43 | 618 | 21 | 578 | ChainA, Beta-galactosidase [Niallia circulans],4MAD_B Chain B, Beta-galactosidase [Niallia circulans] |
| 7KDV_A | 9.26e-119 | 38 | 602 | 17 | 589 | ChainA, Beta-galactosidase [Mus musculus],7KDV_C Chain C, Beta-galactosidase [Mus musculus],7KDV_E Chain E, Beta-galactosidase [Mus musculus],7KDV_G Chain G, Beta-galactosidase [Mus musculus],7KDV_I Chain I, Beta-galactosidase [Mus musculus],7KDV_K Chain K, Beta-galactosidase [Mus musculus] |
| 4E8C_A | 3.28e-115 | 38 | 641 | 3 | 594 | Crystalstructure of streptococcal beta-galactosidase in complex with galactose [Streptococcus pneumoniae TIGR4],4E8C_B Crystal structure of streptococcal beta-galactosidase in complex with galactose [Streptococcus pneumoniae TIGR4],4E8D_A Crystal structure of streptococcal beta-galactosidase [Streptococcus pneumoniae TIGR4],4E8D_B Crystal structure of streptococcal beta-galactosidase [Streptococcus pneumoniae TIGR4] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P48982 | 1.34e-123 | 6 | 600 | 4 | 572 | Beta-galactosidase OS=Xanthomonas manihotis OX=43353 GN=bga PE=1 SV=1 |
| P23780 | 2.27e-117 | 38 | 602 | 34 | 606 | Beta-galactosidase OS=Mus musculus OX=10090 GN=Glb1 PE=1 SV=1 |
| P16278 | 1.09e-110 | 45 | 602 | 40 | 604 | Beta-galactosidase OS=Homo sapiens OX=9606 GN=GLB1 PE=1 SV=2 |
| Q5R7P4 | 2.13e-110 | 46 | 602 | 41 | 604 | Beta-galactosidase OS=Pongo abelii OX=9601 GN=GLB1 PE=2 SV=1 |
| Q95LV1 | 1.83e-108 | 46 | 635 | 39 | 625 | Beta-galactosidase-1-like protein OS=Macaca fascicularis OX=9541 GN=GLB1L PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
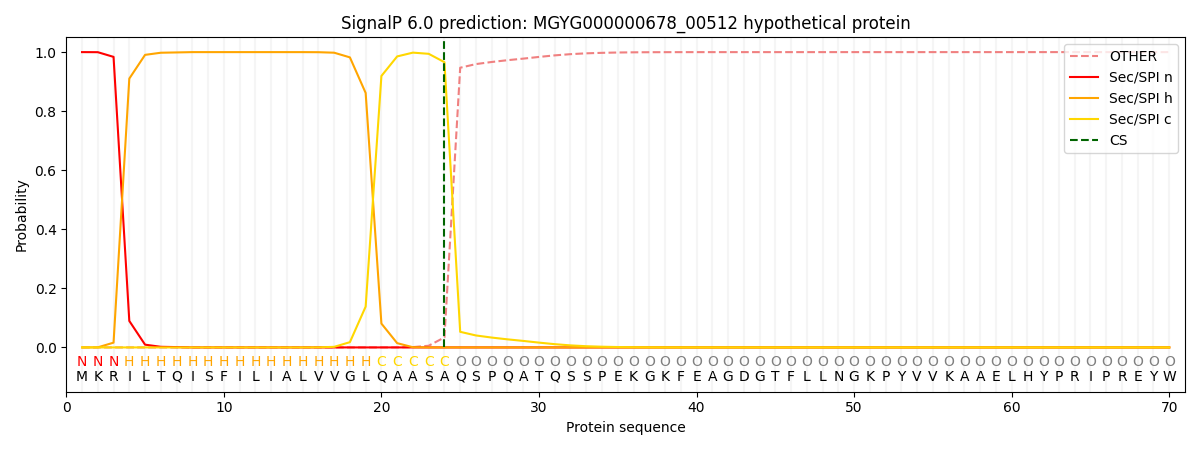
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000305 | 0.998870 | 0.000286 | 0.000180 | 0.000166 | 0.000151 |
