You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000682_01178
You are here: Home > Sequence: MGYG000000682_01178
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; UBA4334; | |||||||||||
| CAZyme ID | MGYG000000682_01178 | |||||||||||
| CAZy Family | GH53 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 18725; End: 20161 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH53 | 41 | 370 | 3.8e-68 | 0.9766081871345029 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07745 | Glyco_hydro_53 | 1.65e-62 | 40 | 370 | 1 | 327 | Glycosyl hydrolase family 53. This domain belongs to family 53 of the glycosyl hydrolase classification. These enzymes are enzymes are endo-1,4- beta-galactanases (EC:3.2.1.89). The structure of this domain is known and has a TIM barrel fold. |
| COG3867 | GanB | 8.28e-40 | 7 | 370 | 5 | 385 | Arabinogalactan endo-1,4-beta-galactosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUB75889.1 | 6.21e-125 | 39 | 454 | 52 | 468 |
| QUB65825.1 | 9.32e-125 | 39 | 454 | 21 | 437 |
| QUB62879.1 | 9.93e-124 | 39 | 454 | 52 | 468 |
| QUB67749.1 | 1.40e-123 | 39 | 454 | 52 | 468 |
| AWB69168.1 | 2.52e-80 | 39 | 465 | 28 | 466 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1R8L_A | 1.60e-22 | 38 | 365 | 23 | 377 | Thestructure of endo-beta-1,4-galactanase from Bacillus licheniformis [Bacillus licheniformis],1R8L_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis [Bacillus licheniformis],1UR0_A The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR0_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR4_A The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR4_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],2CCR_A Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2CCR_B Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2J74_A Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2J74_B Structure of Beta-1,4-Galactanase [Bacillus licheniformis] |
| 2GFT_A | 9.71e-22 | 38 | 365 | 23 | 377 | ChainA, Glycosyl Hydrolase Family 53 [Bacillus licheniformis],2GFT_B Chain B, Glycosyl Hydrolase Family 53 [Bacillus licheniformis] |
| 4BF7_A | 4.73e-21 | 40 | 365 | 21 | 335 | Emericillanidulans endo-beta-1,4-galactanase [Aspergillus nidulans] |
| 1FHL_A | 8.17e-16 | 40 | 365 | 4 | 319 | CrystalStructure Of Beta-1,4-galactanase From Aspergillus Aculeatus At 293k [Aspergillus aculeatus],1FOB_A Crystal Structure Of Beta-1,4-galactanase From Aspergillus Aculeatus At 100k [Aspergillus aculeatus],6Q3R_A ASPERGILLUS ACULEATUS GALACTANASE [Aspergillus aculeatus],6Q3R_B ASPERGILLUS ACULEATUS GALACTANASE [Aspergillus aculeatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P48841 | 1.35e-64 | 39 | 364 | 27 | 351 | Arabinogalactan endo-beta-1,4-galactanase OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=ganB PE=1 SV=1 |
| P48843 | 2.10e-32 | 39 | 370 | 6 | 340 | Uncharacterized protein in bgaB 5'region (Fragment) OS=Niallia circulans OX=1397 PE=3 SV=1 |
| O07013 | 5.15e-24 | 38 | 370 | 52 | 411 | Endo-beta-1,4-galactanase OS=Bacillus subtilis (strain 168) OX=224308 GN=ganB PE=1 SV=1 |
| Q65CX5 | 1.12e-21 | 38 | 365 | 48 | 402 | Endo-beta-1,4-galactanase OS=Bacillus licheniformis (strain ATCC 14580 / DSM 13 / JCM 2505 / CCUG 7422 / NBRC 12200 / NCIMB 9375 / NCTC 10341 / NRRL NRS-1264 / Gibson 46) OX=279010 GN=ganB PE=1 SV=1 |
| Q5B153 | 2.53e-20 | 40 | 365 | 21 | 335 | Arabinogalactan endo-beta-1,4-galactanase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=galA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
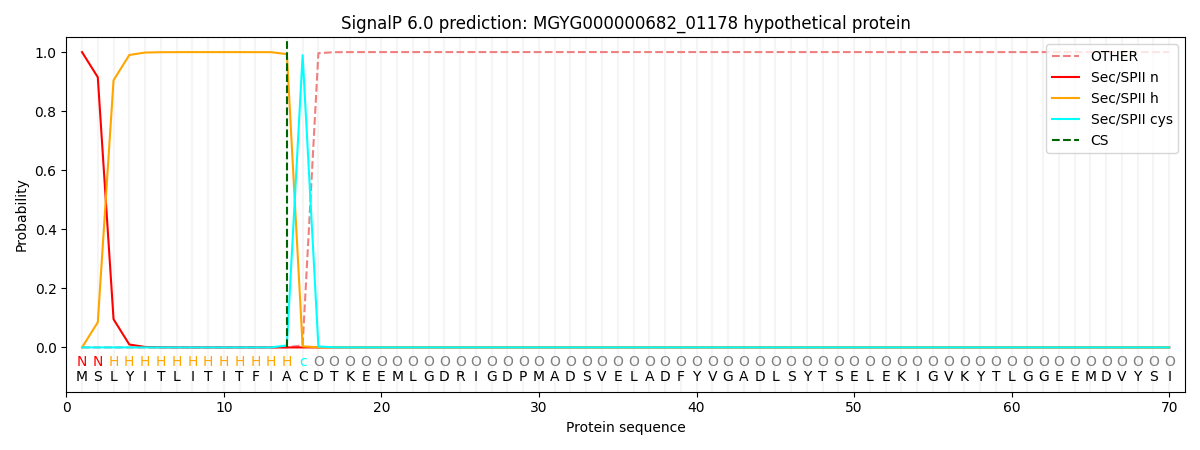
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000027 | 1.000049 | 0.000000 | 0.000000 | 0.000000 |
