You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000709_01336
You are here: Home > Sequence: MGYG000000709_01336
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Oscillospiraceae; UBA738; | |||||||||||
| CAZyme ID | MGYG000000709_01336 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 6; End: 1466 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 1.35e-23 | 2 | 224 | 339 | 565 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| pfam18565 | Glyco_hydro2_C5 | 6.00e-17 | 386 | 475 | 2 | 103 | Glycoside hydrolase family 2 C-terminal domain 5. Domain 5 is found in dimeric beta-D-galactosidase from Paracoccus sp. 32d, which contributes to stabilization of the functional dimer. It is suggested that the location of this domain 5, may be one of the factors responsible for the creation of a functional dimer and cold-adaptation of this enzyme. |
| pfam02836 | Glyco_hydro_2_C | 6.46e-17 | 2 | 219 | 54 | 293 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| PRK10340 | ebgA | 5.05e-13 | 2 | 140 | 373 | 512 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| PRK10150 | PRK10150 | 3.10e-08 | 2 | 90 | 331 | 443 | beta-D-glucuronidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ATO98959.1 | 4.41e-241 | 1 | 482 | 308 | 789 |
| QIA42639.1 | 4.41e-241 | 1 | 482 | 308 | 789 |
| ATL89252.1 | 7.17e-240 | 1 | 482 | 308 | 789 |
| AXB28859.1 | 1.02e-239 | 1 | 482 | 308 | 789 |
| CBL02555.1 | 1.02e-239 | 1 | 482 | 308 | 789 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5EUV_A | 3.54e-95 | 1 | 466 | 298 | 716 | ChainA, Beta-D-galactosidase [Paracoccus sp. 32d],5EUV_B Chain B, Beta-D-galactosidase [Paracoccus sp. 32d],5LDR_B Chain B, Beta-D-galactosidase [Paracoccus sp. 32d] |
| 5LDR_A | 3.62e-95 | 1 | 466 | 299 | 717 | ChainA, Beta-D-galactosidase [Paracoccus sp. 32d] |
| 6NCW_A | 2.84e-18 | 1 | 215 | 308 | 544 | Crystalstructure of a GH2 beta-galacturonidase from Eisenbergiella tayi bound to glycerol [Eisenbergiella tayi],6NCW_B Crystal structure of a GH2 beta-galacturonidase from Eisenbergiella tayi bound to glycerol [Eisenbergiella tayi],6NCW_C Crystal structure of a GH2 beta-galacturonidase from Eisenbergiella tayi bound to glycerol [Eisenbergiella tayi],6NCW_D Crystal structure of a GH2 beta-galacturonidase from Eisenbergiella tayi bound to glycerol [Eisenbergiella tayi],6NCX_A Crystal structure of GH2 beta-galacturonidase from Eisenbergiella tayi bound to galacturonate [Eisenbergiella tayi],6NCX_B Crystal structure of GH2 beta-galacturonidase from Eisenbergiella tayi bound to galacturonate [Eisenbergiella tayi],6NCX_C Crystal structure of GH2 beta-galacturonidase from Eisenbergiella tayi bound to galacturonate [Eisenbergiella tayi],6NCX_D Crystal structure of GH2 beta-galacturonidase from Eisenbergiella tayi bound to galacturonate [Eisenbergiella tayi] |
| 6D1P_A | 8.43e-16 | 1 | 262 | 361 | 657 | Apostructure of Bacteroides uniformis beta-glucuronidase 3 [Bacteroides uniformis str. 3978 T3 ii],6D1P_B Apo structure of Bacteroides uniformis beta-glucuronidase 3 [Bacteroides uniformis str. 3978 T3 ii] |
| 6MVG_A | 7.82e-15 | 1 | 262 | 337 | 615 | Crystalstructure of FMN-binding beta-glucuronidase from Ruminococcus gnavus [[Ruminococcus] gnavus],6MVG_B Crystal structure of FMN-binding beta-glucuronidase from Ruminococcus gnavus [[Ruminococcus] gnavus],6MVG_C Crystal structure of FMN-binding beta-glucuronidase from Ruminococcus gnavus [[Ruminococcus] gnavus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P26257 | 1.27e-104 | 1 | 445 | 322 | 715 | Beta-galactosidase OS=Thermoanaerobacterium thermosulfurigenes OX=33950 GN=lacZ PE=1 SV=1 |
| P77989 | 6.33e-90 | 1 | 483 | 321 | 741 | Beta-galactosidase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=lacZ PE=3 SV=2 |
| Q59750 | 1.79e-53 | 1 | 467 | 314 | 740 | Beta-galactosidase OS=Rhizobium meliloti OX=382 GN=lacZ PE=1 SV=1 |
| T2KPJ7 | 9.57e-20 | 1 | 264 | 363 | 670 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
| T2KM09 | 3.27e-12 | 1 | 262 | 360 | 660 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22050 PE=2 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
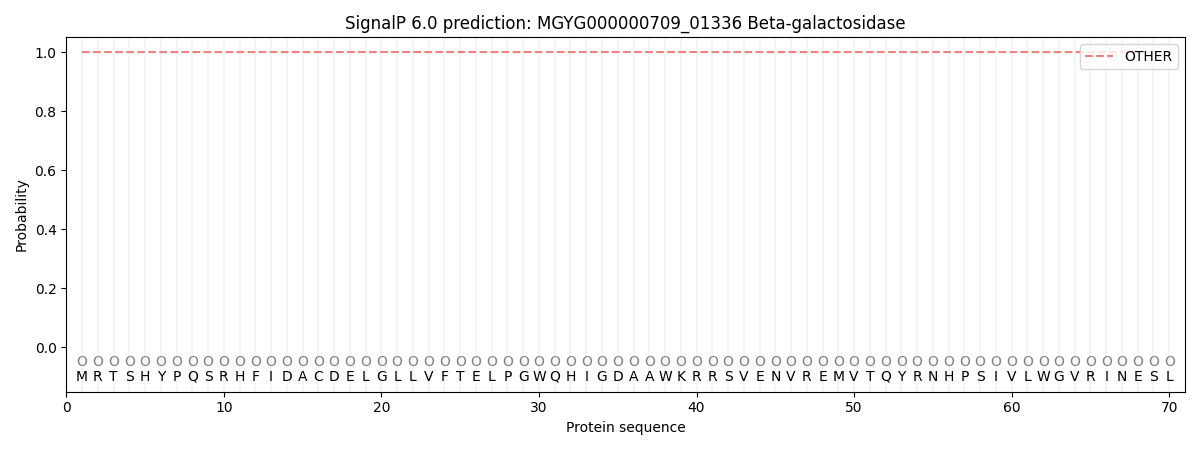
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000068 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
