You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000720_01060
You are here: Home > Sequence: MGYG000000720_01060
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; Victivallaceae; Victivallis; | |||||||||||
| CAZyme ID | MGYG000000720_01060 | |||||||||||
| CAZy Family | GH39 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9220; End: 11082 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH39 | 71 | 252 | 5.6e-28 | 0.5034802784222738 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01229 | Glyco_hydro_39 | 3.62e-09 | 110 | 249 | 122 | 280 | Glycosyl hydrolases family 39. |
| COG3693 | XynA | 0.001 | 6 | 195 | 9 | 218 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
| pfam02449 | Glyco_hydro_42 | 0.001 | 42 | 138 | 10 | 137 | Beta-galactosidase. This group of beta-galactosidase enzymes belong to the glycosyl hydrolase 42 family. The enzyme catalyzes the hydrolysis of terminal, non-reducing terminal beta-D-galactosidase residues. |
| cd00829 | SCP-x_thiolase | 0.002 | 231 | 288 | 221 | 285 | Thiolase domain associated with sterol carrier protein (SCP)-x isoform and related proteins; SCP-2 has multiple roles in intracellular lipid circulation and metabolism. The N-terminal presequence in the SCP-x isoform represents a peroxisomal 3-ketacyl-Coa thiolase specific for branched-chain acyl CoAs, which is proteolytically cleaved from the sterol carrier protein. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM44885.1 | 0.0 | 21 | 616 | 18 | 615 |
| QDU57372.1 | 2.81e-120 | 22 | 619 | 22 | 632 |
| AVM44901.1 | 1.20e-75 | 27 | 531 | 24 | 546 |
| AHF90941.1 | 5.76e-56 | 27 | 554 | 231 | 785 |
| ADE82856.1 | 5.68e-37 | 412 | 617 | 202 | 406 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5JVK_A | 4.16e-13 | 24 | 246 | 102 | 345 | Structuralinsights into a family 39 glycoside hydrolase from the gut symbiont Bacteroides cellulosilyticus WH2. [Bacteroides cellulosilyticus],5JVK_B Structural insights into a family 39 glycoside hydrolase from the gut symbiont Bacteroides cellulosilyticus WH2. [Bacteroides cellulosilyticus],5JVK_C Structural insights into a family 39 glycoside hydrolase from the gut symbiont Bacteroides cellulosilyticus WH2. [Bacteroides cellulosilyticus] |
| 4ZN2_A | 1.84e-12 | 46 | 259 | 37 | 268 | Glycosylhydrolase from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],4ZN2_B Glycosyl hydrolase from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],4ZN2_C Glycosyl hydrolase from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],4ZN2_D Glycosyl hydrolase from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1] |
| 5BX9_A | 1.84e-12 | 46 | 259 | 37 | 268 | Structureof PslG from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5BXA_A Structure of PslG from Pseudomonas aeruginosa in complex with mannose [Pseudomonas aeruginosa PAO1] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
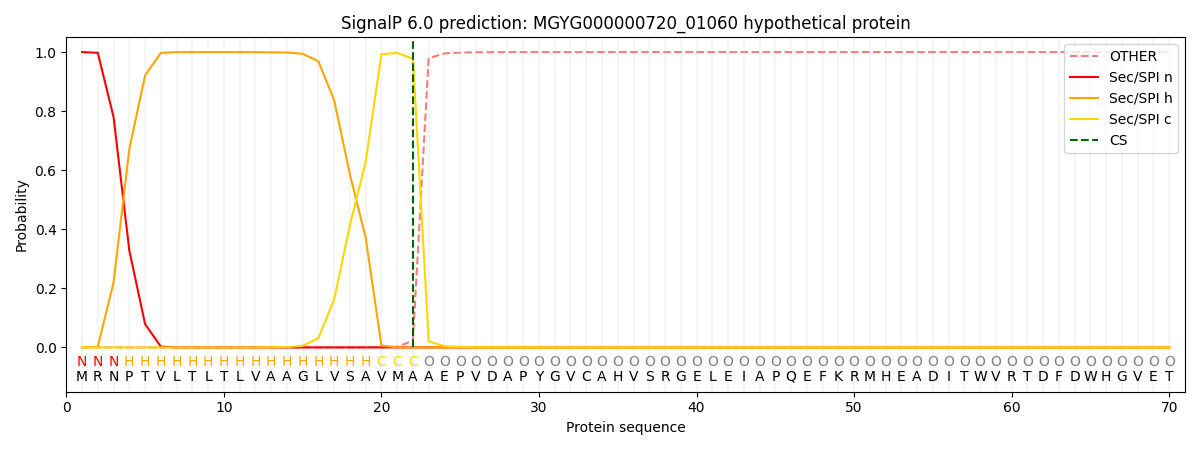
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000200 | 0.999166 | 0.000156 | 0.000169 | 0.000148 | 0.000135 |
