You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000720_01368
You are here: Home > Sequence: MGYG000000720_01368
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; Victivallaceae; Victivallis; | |||||||||||
| CAZyme ID | MGYG000000720_01368 | |||||||||||
| CAZy Family | GH167 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5503; End: 9093 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH167 | 609 | 1187 | 2.2e-120 | 0.8340306834030683 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1874 | GanA | 5.93e-08 | 692 | 1079 | 102 | 490 | Beta-galactosidase GanA [Carbohydrate transport and metabolism]. |
| pfam02449 | Glyco_hydro_42 | 4.99e-07 | 663 | 951 | 30 | 342 | Beta-galactosidase. This group of beta-galactosidase enzymes belong to the glycosyl hydrolase 42 family. The enzyme catalyzes the hydrolysis of terminal, non-reducing terminal beta-D-galactosidase residues. |
| cd09621 | CBM9_like_5 | 7.22e-07 | 231 | 379 | 25 | 179 | DOMON-like type 9 carbohydrate binding module. Family 9 carbohydrate-binding modules (CBM9) play a role in the microbial degradation of cellulose and hemicellulose (materials found in plants). The domain has previously been called cellulose-binding domain. The polysaccharide binding sites of CBMs with available 3D structure have been found to be either flat surfaces with interactions formed by predominantly aromatic residues (tryptophan and tyrosine), or extended shallow grooves. CBM9 domains found in this uncharacterized heterogeneous subfamily are often located at the C-terminus of longer proteins and may co-occur with various other functional domains such as glycosyl hydrolases. The CBM9 module in these architectures may be involved in binding to carbohydrates. |
| cd03143 | A4_beta-galactosidase_middle_domain | 0.004 | 1048 | 1096 | 55 | 103 | A4 beta-galactosidase middle domain: a type 1 glutamine amidotransferase (GATase1)-like domain. A4 beta-galactosidase middle domain: a type 1 glutamine amidotransferase (GATase1)-like domain. This group includes proteins similar to beta-galactosidase from Thermus thermophilus. Beta-Galactosidase hydrolyzes the beta-1,4-D-galactosidic linkage of lactose, as well as those of related chromogens, o-nitrophenyl-beta-D-galactopyranoside (ONP-Gal) and 5-bromo-4-chloro-3-indolyl-beta-D-galactoside (X-gal). This A4 beta-galactosidase middle domain lacks the catalytic triad of typical GATase1 domains. The reactive Cys residue found in the sharp turn between a beta strand and an alpha helix termed the nucleophile elbow in typical GATase1 domains is not conserved in this group. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM43323.1 | 8.21e-217 | 30 | 1189 | 183 | 1356 |
| AVM46829.1 | 4.14e-201 | 4 | 1189 | 1 | 1180 |
| AVM43886.1 | 5.03e-133 | 8 | 1185 | 3 | 994 |
| AHF94331.1 | 7.82e-92 | 58 | 1188 | 218 | 1387 |
| QHW30800.1 | 1.70e-81 | 209 | 1166 | 498 | 1438 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
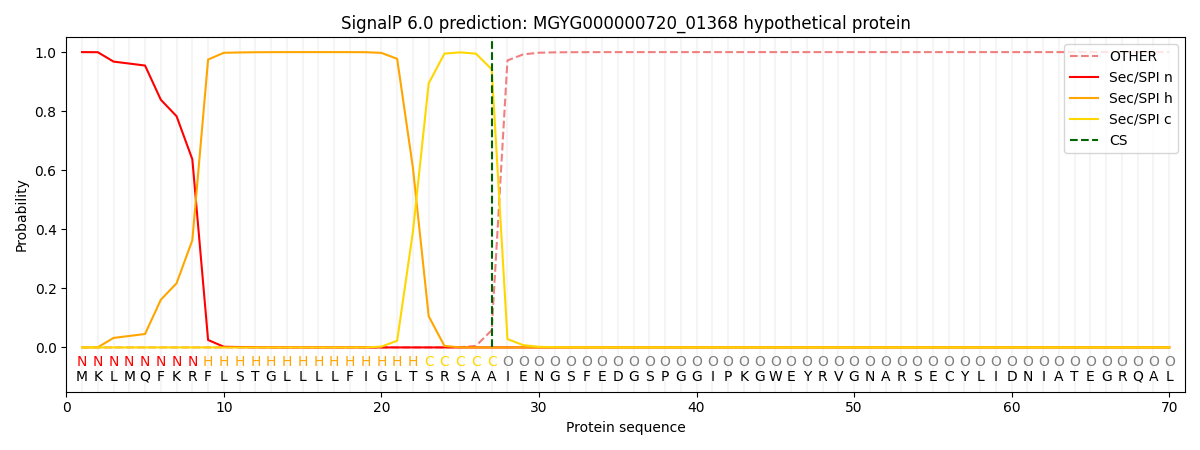
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000254 | 0.999045 | 0.000223 | 0.000162 | 0.000152 | 0.000139 |
