You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000745_01123
You are here: Home > Sequence: MGYG000000745_01123
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; CAG-74; ; | |||||||||||
| CAZyme ID | MGYG000000745_01123 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 29759; End: 31402 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 77 | 338 | 3.1e-106 | 0.9923954372623575 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 6.05e-27 | 100 | 337 | 27 | 268 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 5.73e-21 | 95 | 234 | 71 | 214 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| COG3934 | COG3934 | 7.58e-04 | 101 | 368 | 30 | 313 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
| cd09755 | Cas2_I-E | 0.005 | 190 | 229 | 17 | 56 | CRISPR/Cas system-associated protein Cas2. CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) and associated Cas proteins comprise a system for heritable host defense by prokaryotic cells against phage and other foreign DNA; Cas2 is present in majority of CRISPR/Cas systems along with Cas1; RNAse specific to U-rich regions; Possesses an RRM/ferredoxin fold |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| VEI36204.1 | 1.79e-122 | 23 | 440 | 36 | 447 |
| AZS15177.1 | 4.31e-121 | 30 | 373 | 49 | 396 |
| AIC93076.1 | 3.19e-112 | 28 | 385 | 29 | 393 |
| AQR93158.1 | 1.90e-92 | 28 | 369 | 919 | 1263 |
| AMK77044.1 | 4.11e-89 | 30 | 375 | 35 | 394 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1H4P_A | 3.91e-21 | 47 | 274 | 13 | 258 | Crystalstructure of exo-1,3-beta glucanse from Saccharomyces cerevisiae [Saccharomyces cerevisiae],1H4P_B Crystal structure of exo-1,3-beta glucanse from Saccharomyces cerevisiae [Saccharomyces cerevisiae] |
| 3O6A_A | 1.72e-18 | 34 | 274 | 1 | 255 | F144Y/F258YDouble Mutant of Exo-beta-1,3-glucanase from Candida albicans at 2 A [Candida albicans] |
| 1EQP_A | 2.95e-18 | 41 | 274 | 3 | 250 | Exo-b-(1,3)-glucanaseFrom Candida Albicans [Candida albicans] |
| 4M80_A | 3.08e-18 | 34 | 274 | 1 | 255 | Thestructure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans at 1.85A resolution [Candida albicans SC5314],4M81_A The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans complexed with 1-fluoro-alpha-D-glucopyranoside (donor) and p-nitrophenyl beta-D-glucopyranoside (acceptor) at 1.86A resolution [Candida albicans SC5314],4M82_A The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans complexed with p-nitrophenyl-gentiobioside (product) at 1.6A resolution [Candida albicans SC5314] |
| 3N9K_A | 3.08e-18 | 34 | 274 | 1 | 255 | F229A/E292SDouble Mutant of Exo-beta-1,3-glucanase from Candida albicans in Complex with Laminaritriose at 1.7 A [Candida albicans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| W8QRE4 | 1.24e-38 | 30 | 273 | 4 | 248 | Beta-xylosidase OS=Phanerodontia chrysosporium OX=2822231 GN=Xyl5 PE=1 SV=2 |
| A2RAR6 | 6.89e-25 | 21 | 252 | 17 | 250 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=exgA PE=3 SV=1 |
| A1D4Q5 | 1.87e-23 | 46 | 248 | 41 | 247 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=exgA PE=3 SV=1 |
| B0XN12 | 3.40e-23 | 46 | 248 | 41 | 247 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=exgA PE=3 SV=1 |
| B8N151 | 4.05e-23 | 41 | 213 | 27 | 200 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=exgA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
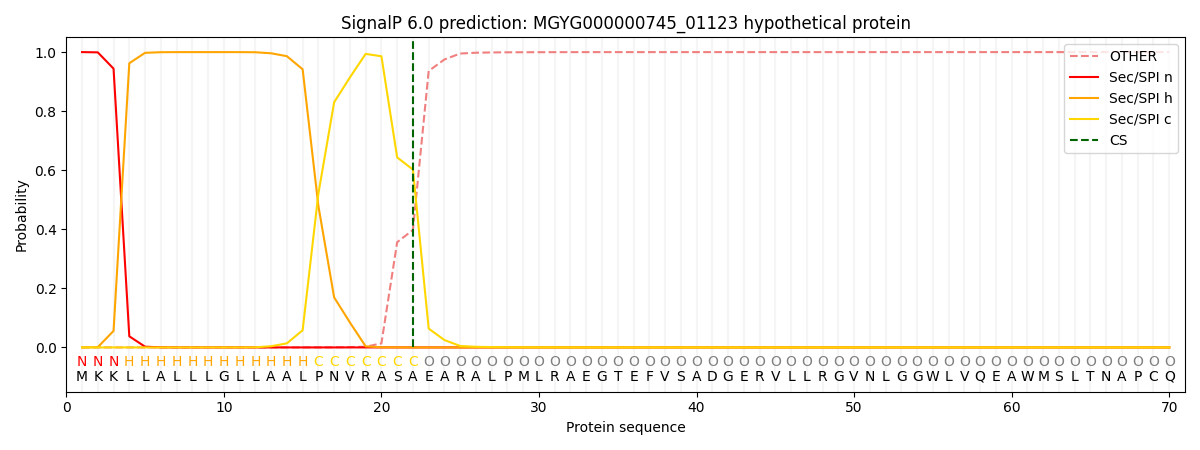
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000282 | 0.999046 | 0.000163 | 0.000183 | 0.000159 | 0.000148 |
