You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000747_01539
You are here: Home > Sequence: MGYG000000747_01539
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; CAG-74; ; | |||||||||||
| CAZyme ID | MGYG000000747_01539 | |||||||||||
| CAZy Family | GH36 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3976; End: 5586 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd14791 | GH36 | 2.03e-29 | 195 | 489 | 2 | 296 | glycosyl hydrolase family 36 (GH36). GH36 enzymes occur in prokaryotes, eukaryotes, and archaea with a wide range of hydrolytic activities, including alpha-galactosidase, alpha-N-acetylgalactosaminidase, stachyose synthase, and raffinose synthase. All GH36 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH36 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
| cd14790 | GH_D | 7.03e-05 | 228 | 455 | 29 | 219 | Glycoside hydrolases, clan D. This group of glycosyl hydrolase families is comprised of glycosyl hydrolase family 31 (GH31), family 36 (GH36), and family 27 (GH27). These structurally and mechanistically related protein families are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. They have a wide range of functions including alpha-glucosidase, alpha-xylosidase, 6-alpha-glucosyltransferase, 3-alpha-isomaltosyltransferase, alpha-N-acetylgalactosaminidase, stachyose synthase, raffinose synthase, and alpha-1,4-glucan lyase. |
| COG3345 | GalA | 2.32e-04 | 198 | 444 | 295 | 568 | Alpha-galactosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| SNU96820.1 | 3.01e-169 | 1 | 523 | 6 | 523 |
| CBL19648.1 | 1.45e-168 | 5 | 530 | 30 | 547 |
| QCU01578.1 | 4.13e-168 | 5 | 530 | 30 | 547 |
| BDA09953.1 | 3.68e-163 | 1 | 536 | 6 | 538 |
| QIB59833.1 | 1.48e-162 | 1 | 523 | 6 | 526 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
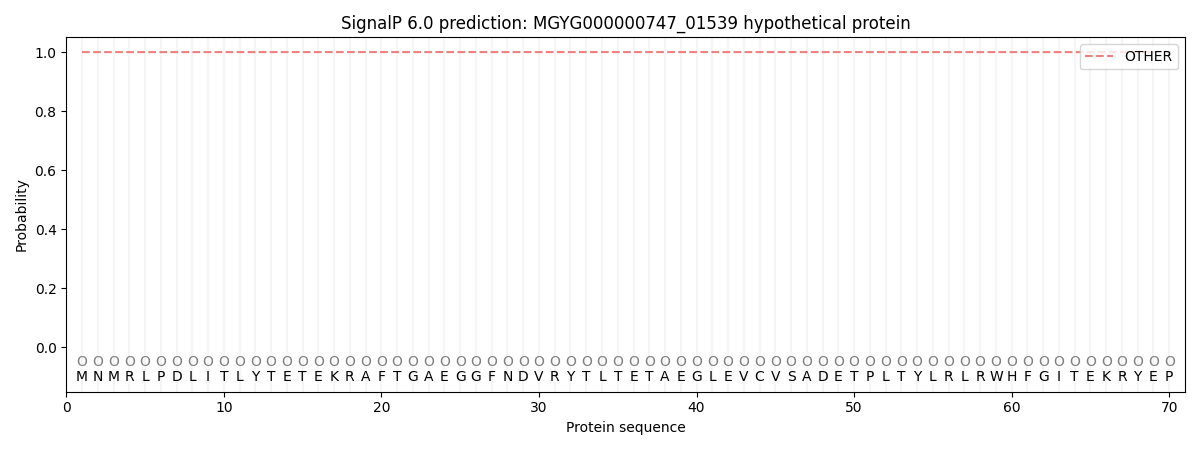
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000050 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
