You are browsing environment: HUMAN GUT
MGYG000000752_01102
Basic Information
help
Species
Lineage
Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; UBA737;
CAZyme ID
MGYG000000752_01102
CAZy Family
GH163
CAZyme Description
hypothetical protein
CAZyme Property
Protein Length
CGC
Molecular Weight
Isoelectric Point
585
68111.18
5.7971
Genome Property
Genome Assembly ID
Genome Size
Genome Type
Country
Continent
MGYG000000752
2697118
MAG
Kazakhstan
Asia
Gene Location
Start: 15789;
End: 17546
Strand: -
No EC number prediction in MGYG000000752_01102.
Family
Start
End
Evalue
family coverage
GH163
197
460
9.2e-90
0.9920318725099602
Cdd ID
Domain
E-Value
qStart
qEnd
sStart
sEnd
Domain Description
pfam16126
DUF4838
1.70e-102
189
460
1
263
Domain of unknown function (DUF4838). This family consists of several uncharacterized proteins found in various Bacteroides and Chloroflexus species. The function of this family is unknown.
more
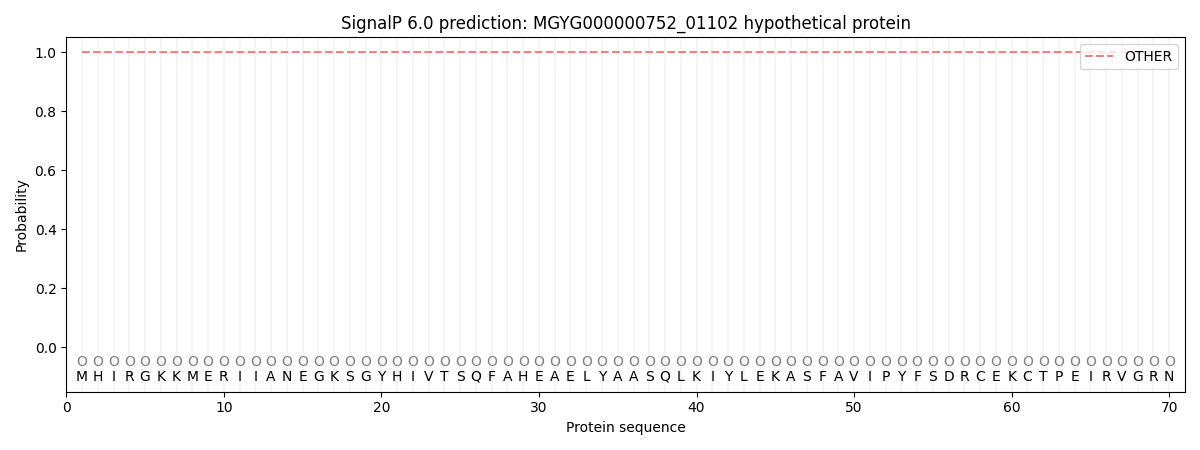
This protein is predicted as OTHER
Other
SP_Sec_SPI
LIPO_Sec_SPII
TAT_Tat_SPI
TATLIP_Sec_SPII
PILIN_Sec_SPIII
1.000057
0.000000
0.000000
0.000000
0.000000
0.000000
There is no transmembrane helices in MGYG000000752_01102.