You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000765_01362
You are here: Home > Sequence: MGYG000000765_01362
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Porphyromonas_A bennonis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Porphyromonadaceae; Porphyromonas_A; Porphyromonas_A bennonis | |||||||||||
| CAZyme ID | MGYG000000765_01362 | |||||||||||
| CAZy Family | GH29 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 16482; End: 18314 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH29 | 57 | 378 | 3.6e-80 | 0.8728323699421965 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3669 | AfuC | 8.73e-70 | 57 | 469 | 11 | 417 | Alpha-L-fucosidase [Carbohydrate transport and metabolism]. |
| pfam01120 | Alpha_L_fucos | 7.78e-40 | 95 | 380 | 79 | 333 | Alpha-L-fucosidase. |
| smart00812 | Alpha_L_fucos | 4.53e-32 | 64 | 376 | 23 | 331 | Alpha-L-fucosidase. O-Glycosyl hydrolases (EC 3.2.1.-) are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycosyl hydrolases, based on sequence similarity, has led to the definition of 85 different families. This classification is available on the CAZy (CArbohydrate-Active EnZymes) web site. Because the fold of proteins is better conserved than their sequences, some of the families can be grouped in 'clans'. Family 29 encompasses alpha-L-fucosidases, which is a lysosomal enzyme responsible for hydrolyzing the alpha-1,6-linked fucose joined to the reducing-end N-acetylglucosamine of the carbohydrate moieties of glycoproteins. Deficiency of alpha-L-fucosidase results in the lysosomal storage disease fucosidosis. |
| pfam00754 | F5_F8_type_C | 1.08e-04 | 507 | 594 | 28 | 114 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT71943.1 | 2.41e-213 | 26 | 607 | 21 | 600 |
| QNL38600.1 | 3.31e-213 | 19 | 594 | 13 | 586 |
| QGT71427.1 | 4.69e-213 | 20 | 594 | 14 | 586 |
| QUT80690.1 | 4.69e-213 | 20 | 594 | 14 | 586 |
| QDM09076.1 | 4.69e-213 | 20 | 594 | 14 | 586 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4ZRX_A | 7.14e-209 | 26 | 607 | 1 | 580 | Crystalstructure of a putative alpha-L-fucosidase (BACOVA_04357) from Bacteroides ovatus ATCC 8483 at 1.59 A resolution [Bacteroides ovatus ATCC 8483] |
| 3UES_A | 1.10e-131 | 41 | 471 | 3 | 465 | Crystalstructure of alpha-1,3/4-fucosidase from Bifidobacterium longum subsp. infantis complexed with deoxyfuconojirimycin [Bifidobacterium longum subsp. infantis ATCC 15697 = JCM 1222 = DSM 20088],3UES_B Crystal structure of alpha-1,3/4-fucosidase from Bifidobacterium longum subsp. infantis complexed with deoxyfuconojirimycin [Bifidobacterium longum subsp. infantis ATCC 15697 = JCM 1222 = DSM 20088] |
| 3MO4_A | 7.44e-130 | 41 | 471 | 5 | 467 | Thecrystal structure of an alpha-(1-3,4)-fucosidase from Bifidobacterium longum subsp. infantis ATCC 15697 [Bifidobacterium longum subsp. infantis ATCC 15697 = JCM 1222 = DSM 20088],3MO4_B The crystal structure of an alpha-(1-3,4)-fucosidase from Bifidobacterium longum subsp. infantis ATCC 15697 [Bifidobacterium longum subsp. infantis ATCC 15697 = JCM 1222 = DSM 20088] |
| 3UET_A | 1.39e-129 | 41 | 471 | 3 | 465 | Crystalstructure of alpha-1,3/4-fucosidase from Bifidobacterium longum subsp. infantis D172A/E217A mutant complexed with lacto-N-fucopentaose II [Bifidobacterium longum subsp. infantis ATCC 15697 = JCM 1222 = DSM 20088],3UET_B Crystal structure of alpha-1,3/4-fucosidase from Bifidobacterium longum subsp. infantis D172A/E217A mutant complexed with lacto-N-fucopentaose II [Bifidobacterium longum subsp. infantis ATCC 15697 = JCM 1222 = DSM 20088] |
| 5K9H_A | 7.10e-102 | 29 | 578 | 12 | 554 | Crystalstructure of a glycoside hydrolase 29 family member from an unknown rumen bacterium [unidentified] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8GW72 | 2.95e-111 | 56 | 471 | 37 | 466 | Alpha-L-fucosidase 1 OS=Arabidopsis thaliana OX=3702 GN=FUC1 PE=1 SV=2 |
| Q7XUR3 | 1.64e-99 | 56 | 471 | 39 | 465 | Putative alpha-L-fucosidase 1 OS=Oryza sativa subsp. japonica OX=39947 GN=Os04g0560400 PE=3 SV=2 |
| P49713 | 1.94e-09 | 95 | 399 | 92 | 366 | Putative alpha-L-fucosidase OS=Caenorhabditis elegans OX=6239 GN=W03G11.3 PE=3 SV=2 |
| Q99KR8 | 5.66e-09 | 64 | 376 | 72 | 355 | Plasma alpha-L-fucosidase OS=Mus musculus OX=10090 GN=Fuca2 PE=1 SV=1 |
| Q9BTY2 | 7.60e-09 | 64 | 376 | 78 | 361 | Plasma alpha-L-fucosidase OS=Homo sapiens OX=9606 GN=FUCA2 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
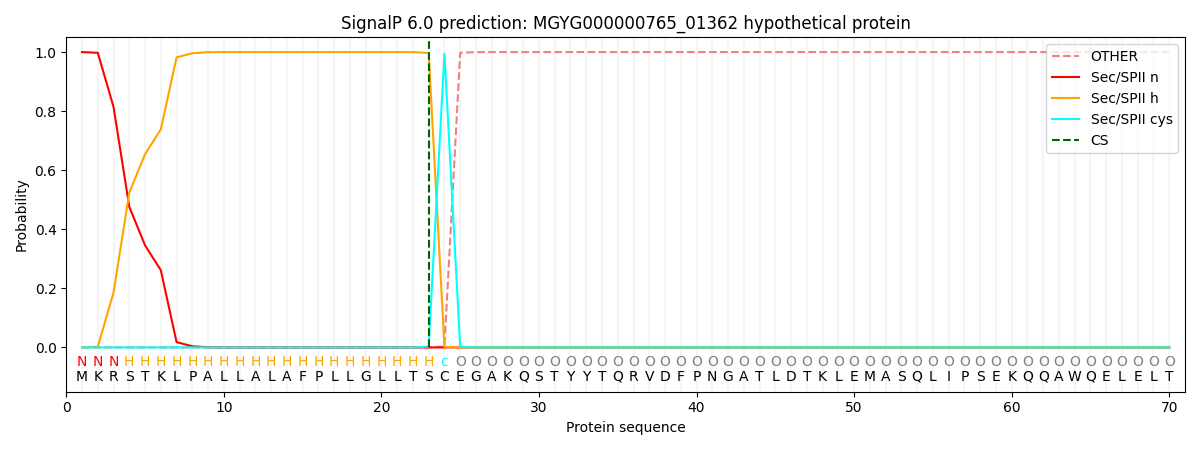
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000086 | 0.000000 | 0.000000 | 0.000000 |
