You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000779_00530
You are here: Home > Sequence: MGYG000000779_00530
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp000431975 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp000431975 | |||||||||||
| CAZyme ID | MGYG000000779_00530 | |||||||||||
| CAZy Family | GH109 | |||||||||||
| CAZyme Description | Glycosyl hydrolase family 109 protein 1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 100; End: 1809 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH109 | 63 | 473 | 5.5e-158 | 0.9899749373433584 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0673 | MviM | 2.79e-20 | 64 | 393 | 1 | 303 | Predicted dehydrogenase [General function prediction only]. |
| pfam01408 | GFO_IDH_MocA | 2.63e-11 | 67 | 192 | 1 | 118 | Oxidoreductase family, NAD-binding Rossmann fold. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
| PRK11579 | PRK11579 | 1.33e-07 | 67 | 188 | 5 | 116 | putative oxidoreductase; Provisional |
| PRK10206 | PRK10206 | 6.27e-04 | 135 | 185 | 63 | 113 | putative oxidoreductase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRO24012.1 | 0.0 | 1 | 512 | 1 | 513 |
| QCQ31109.1 | 0.0 | 6 | 510 | 7 | 511 |
| QPH58140.1 | 2.13e-305 | 8 | 513 | 14 | 512 |
| QUT62233.1 | 3.03e-305 | 8 | 513 | 14 | 512 |
| QQA30686.1 | 3.03e-305 | 8 | 513 | 14 | 512 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6T2B_A | 3.40e-85 | 49 | 472 | 25 | 438 | Glycosidehydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_B Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_C Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_D Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila] |
| 2IXA_A | 1.71e-64 | 65 | 476 | 19 | 435 | A-zyme,N-acetylgalactosaminidase [Elizabethkingia meningoseptica],2IXB_A Crystal structure of N-ACETYLGALACTOSAMINIDASE in complex with GalNAC [Elizabethkingia meningoseptica] |
| 3E82_A | 4.56e-07 | 66 | 273 | 12 | 191 | Crystalstructure of a putative oxidoreductase from Klebsiella pneumoniae [Micrococcus luteus NCTC 2665],3E82_B Crystal structure of a putative oxidoreductase from Klebsiella pneumoniae [Micrococcus luteus NCTC 2665],3E82_D Crystal structure of a putative oxidoreductase from Klebsiella pneumoniae [Micrococcus luteus NCTC 2665],3E82_E Crystal structure of a putative oxidoreductase from Klebsiella pneumoniae [Micrococcus luteus NCTC 2665] |
| 3E9M_A | 9.11e-06 | 103 | 234 | 39 | 167 | CrystalStructure of an oxidoreductase from Enterococcus faecalis [Enterococcus faecalis],3E9M_B Crystal Structure of an oxidoreductase from Enterococcus faecalis [Enterococcus faecalis],3E9M_C Crystal Structure of an oxidoreductase from Enterococcus faecalis [Enterococcus faecalis],3E9M_D Crystal Structure of an oxidoreductase from Enterococcus faecalis [Enterococcus faecalis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A6L1Z2 | 2.22e-303 | 1 | 516 | 1 | 517 | Glycosyl hydrolase family 109 protein 5 OS=Phocaeicola vulgatus (strain ATCC 8482 / DSM 1447 / JCM 5826 / CCUG 4940 / NBRC 14291 / NCTC 11154) OX=435590 GN=BVU_2041 PE=3 SV=1 |
| A6KWM1 | 9.50e-297 | 7 | 516 | 8 | 516 | Glycosyl hydrolase family 109 protein 4 OS=Phocaeicola vulgatus (strain ATCC 8482 / DSM 1447 / JCM 5826 / CCUG 4940 / NBRC 14291 / NCTC 11154) OX=435590 GN=BVU_0105 PE=3 SV=1 |
| A6KYY1 | 1.06e-244 | 6 | 503 | 5 | 498 | Glycosyl hydrolase family 109 protein 3 OS=Phocaeicola vulgatus (strain ATCC 8482 / DSM 1447 / JCM 5826 / CCUG 4940 / NBRC 14291 / NCTC 11154) OX=435590 GN=BVU_0950 PE=3 SV=1 |
| A6KX96 | 6.73e-212 | 44 | 483 | 38 | 470 | Glycosyl hydrolase family 109 protein 1 OS=Phocaeicola vulgatus (strain ATCC 8482 / DSM 1447 / JCM 5826 / CCUG 4940 / NBRC 14291 / NCTC 11154) OX=435590 GN=BVU_0340 PE=3 SV=1 |
| Q5LGZ0 | 6.97e-210 | 2 | 483 | 3 | 463 | Glycosyl hydrolase family 109 protein 1 OS=Bacteroides fragilis (strain ATCC 25285 / DSM 2151 / CCUG 4856 / JCM 11019 / NCTC 9343 / Onslow) OX=272559 GN=BF0853 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
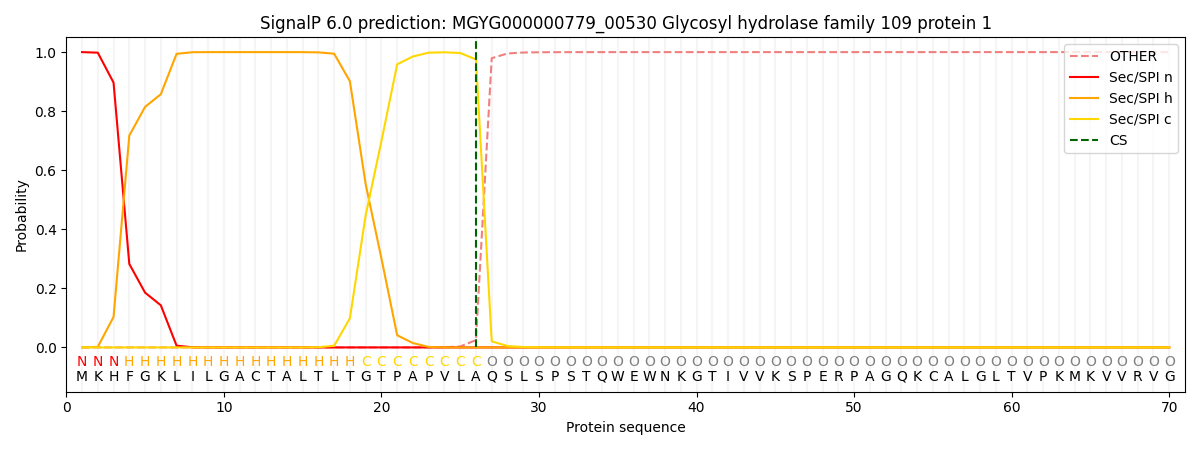
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000286 | 0.998999 | 0.000202 | 0.000188 | 0.000159 | 0.000138 |
