You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000793_00665
You are here: Home > Sequence: MGYG000000793_00665
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
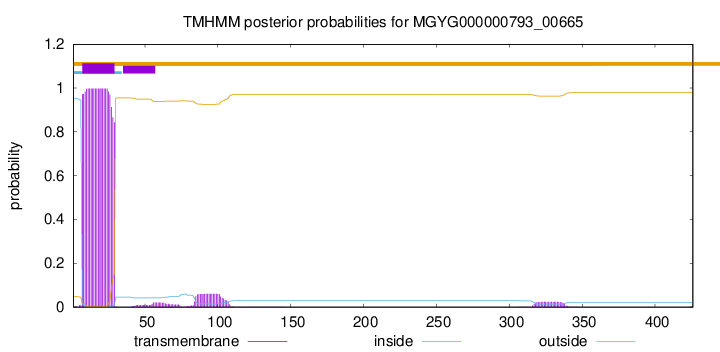
TMHMM annotations
Basic Information help
| Species | Succinivibrio sp900548485 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Succinivibrionaceae; Succinivibrio; Succinivibrio sp900548485 | |||||||||||
| CAZyme ID | MGYG000000793_00665 | |||||||||||
| CAZy Family | GT30 | |||||||||||
| CAZyme Description | 3-deoxy-D-manno-octulosonic acid transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4364; End: 5644 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT30 | 42 | 215 | 6.3e-54 | 0.9717514124293786 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK05749 | PRK05749 | 7.19e-132 | 10 | 424 | 4 | 421 | 3-deoxy-D-manno-octulosonic-acid transferase; Reviewed |
| COG1519 | KdtA | 5.58e-120 | 9 | 422 | 2 | 417 | 3-deoxy-D-manno-octulosonic-acid transferase [Cell wall/membrane/envelope biogenesis]. |
| pfam04413 | Glycos_transf_N | 4.08e-48 | 42 | 215 | 1 | 176 | 3-Deoxy-D-manno-octulosonic-acid transferase (kdotransferase). Members of this family transfer activated sugars to a variety of substrates, including glycogen, fructose-6-phosphate and lipopolysaccharides. Members of the family transfer UDP, ADP, GDP or CMP linked sugars. The Glycos_transf_N region is flanked at the N-terminus by a signal peptide and at the C-terminus by Glycos_transf_1 (pfam00534). The eukaryotic glycogen synthases may be distant members of this bacterial family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AUD79454.1 | 3.13e-83 | 10 | 424 | 4 | 415 |
| ACQ91766.1 | 2.11e-79 | 11 | 422 | 21 | 432 |
| AHN27034.1 | 4.67e-78 | 11 | 417 | 4 | 413 |
| QFT34959.1 | 5.29e-77 | 11 | 420 | 5 | 419 |
| QGN68360.1 | 5.29e-77 | 11 | 420 | 5 | 419 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2XCI_A | 1.47e-44 | 42 | 397 | 24 | 352 | Membrane-embeddedmonofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_B Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_C Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_D Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCU_A Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_B Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_C Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_D Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus] |
| 4BFC_A | 4.01e-18 | 233 | 415 | 41 | 222 | ChainA, 3-DEOXY-D-MANNO-OCTULOSONIC-ACID TRANSFERASE [Acinetobacter baumannii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P44806 | 1.53e-74 | 10 | 417 | 4 | 416 | 3-deoxy-D-manno-octulosonic acid transferase OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=waaA PE=1 SV=1 |
| P0AC75 | 1.72e-64 | 11 | 418 | 5 | 414 | 3-deoxy-D-manno-octulosonic acid transferase OS=Escherichia coli (strain K12) OX=83333 GN=waaA PE=1 SV=1 |
| P0AC77 | 1.72e-64 | 11 | 418 | 5 | 414 | 3-deoxy-D-manno-octulosonic acid transferase OS=Escherichia coli O157:H7 OX=83334 GN=waaA PE=3 SV=1 |
| P0AC76 | 1.72e-64 | 11 | 418 | 5 | 414 | 3-deoxy-D-manno-octulosonic acid transferase OS=Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC) OX=199310 GN=waaA PE=3 SV=1 |
| Q8R6G8 | 6.53e-52 | 30 | 391 | 18 | 377 | Bifunctional glycosyltransferase/methyltransferase OS=Fusobacterium nucleatum subsp. nucleatum (strain ATCC 25586 / DSM 15643 / BCRC 10681 / CIP 101130 / JCM 8532 / KCTC 2640 / LMG 13131 / VPI 4355) OX=190304 GN=trmB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
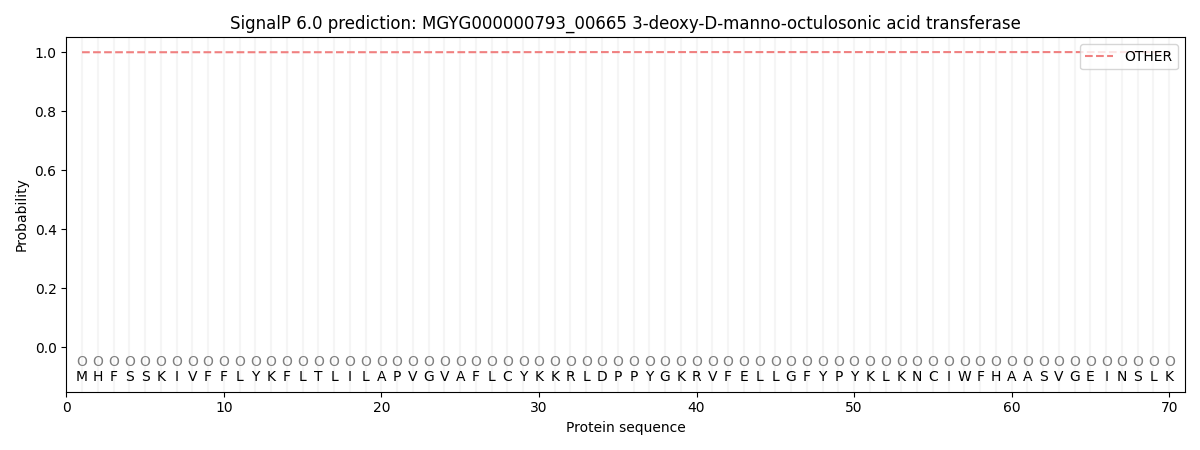
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999570 | 0.000457 | 0.000002 | 0.000001 | 0.000001 | 0.000003 |