You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000820_00083
You are here: Home > Sequence: MGYG000000820_00083
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA9506 sp003506415 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales_A; UBA9506; UBA9506; UBA9506 sp003506415 | |||||||||||
| CAZyme ID | MGYG000000820_00083 | |||||||||||
| CAZy Family | GH57 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 77272; End: 78813 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH57 | 5 | 349 | 1.5e-31 | 0.7963446475195822 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10794 | GH57N_PfGalA_like | 3.03e-35 | 4 | 159 | 1 | 153 | N-terminal catalytic domain of alpha-galactosidase; glycoside hydrolase family 57 (GH57). Alpha-galactosidases (GalA, EC 3.2.1.22) catalyze the hydrolysis of alpha-1,6-linked galactose residues from oligosaccharides and polymeric galactomannans. Based on sequence similarity, the majority of eukaryotic and bacterial GalAs have been classified into glycoside hydrolase family GH27, GH36, and GH4, respectively. This subfamily is represented by a novel type of GalA from Pyrococcus furiosus (PfGalA), which belongs to the GH57 family. PfGalA is an extremely thermo-active and thermostable GalA that functions as a bacterial-like GalA, however, without the capacity to hydrolyze polysaccharides. It specifically catalyzes the hydrolysis of para-nitrophenyl-alpha-galactopyranoside, and to some extent that of melibiose and raffinose. PfGalA has a pH optimum between 5.0-5.5. |
| pfam03065 | Glyco_hydro_57 | 4.93e-21 | 6 | 220 | 23 | 252 | Glycosyl hydrolase family 57. This family includes alpha-amylase (EC:3.2.1.1), 4--glucanotransferase (EC:2.4.1.-) and amylopullulanase enzymes. |
| cd01022 | GH57N_like | 5.68e-18 | 6 | 223 | 3 | 247 | N-terminal catalytic domain of heat stable retaining glycoside hydrolase family 57. Glycoside hydrolase family 57(GH57) is a chiefly prokaryotic family with the majority of thermostable enzymes coming from extremophiles (many of these are archaeal hyperthermophiles), which exhibit the enzyme specificities of alpha-amylase (EC 3.2.1.1), 4-alpha-glucanotransferase (EC 2.4.1.25), amylopullulanase (EC 3.2.1.1/41), and alpha-galactosidase (EC 3.2.1.22). This family also includes many hypothetical proteins with uncharacterized activity and specificity. GH57s cleave alpha-glycosidic bonds by employing a retaining mechanism, which involves a glycosyl-enzyme intermediate, allowing transglycosylation. |
| cd10793 | GH57N_TLGT_like | 2.51e-14 | 21 | 138 | 19 | 135 | N-terminal catalytic domain of 4-alpha-glucanotransferase; glycoside hydrolase family 57 (GH57). 4-alpha-glucanotransferase (TLGT, EC 2.4.1.25) plays a key role in the maltose metabolism. It catalyzes the disproportionation of amylose and the formation of large cyclic alpha-1,4-glucan (cycloamylose) from linear amylose. TLGT functions as a homodimer. Each monomer is composed of two domains, an N-terminal catalytic domain with a (beta/alpha)7 barrel fold and a C-terminal domain with a twisted beta-sandwich fold. Some family members have been designated as alpha-amylases, such as the heat-stable eubacterial amylase from Dictyoglomus thermophilum (DtAmyA) and the extremely thermostable archaeal amylase from Pyrococcus furiosus(PfAmyA). However, both of these proteins are 4-alpha-glucanotransferases. DtAmyA was shown to have transglycosylating activity and PfAmyA exhibits 4-alpha-glucanotransferase activity. |
| COG1449 | COG1449 | 2.24e-12 | 36 | 487 | 152 | 549 | Alpha-amylase/alpha-mannosidase, GH57 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDT00377.1 | 8.69e-254 | 1 | 513 | 1 | 511 |
| AVM45997.1 | 3.70e-44 | 8 | 446 | 12 | 455 |
| AHF90539.1 | 3.54e-42 | 8 | 419 | 11 | 426 |
| EHR78079.1 | 1.72e-16 | 5 | 168 | 4 | 167 |
| CAT70989.1 | 5.73e-15 | 5 | 178 | 4 | 177 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
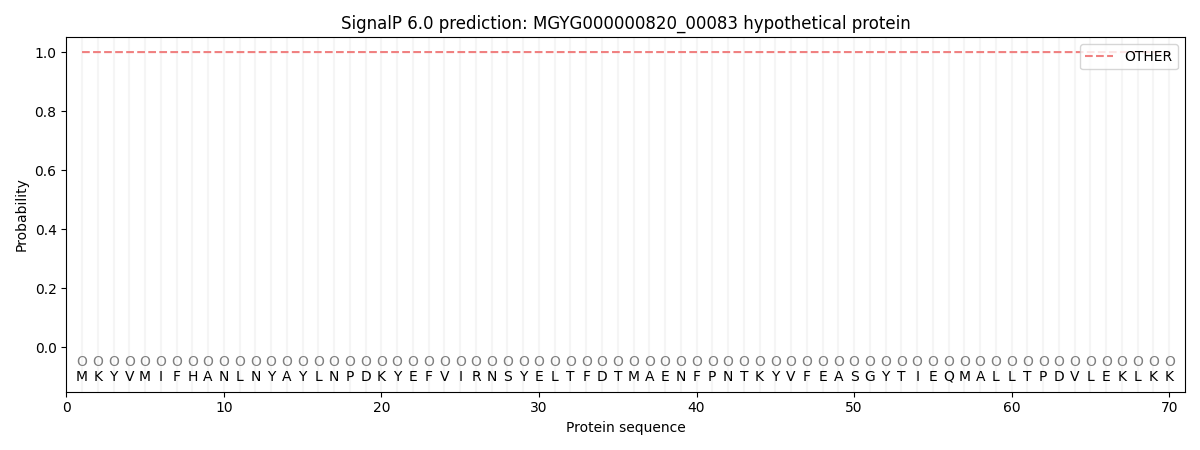
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000075 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
