You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000820_01508
You are here: Home > Sequence: MGYG000000820_01508
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA9506 sp003506415 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales_A; UBA9506; UBA9506; UBA9506 sp003506415 | |||||||||||
| CAZyme ID | MGYG000000820_01508 | |||||||||||
| CAZy Family | GH39 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 86194; End: 87549 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH39 | 71 | 407 | 2.5e-31 | 0.7470997679814385 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01229 | Glyco_hydro_39 | 5.55e-12 | 77 | 207 | 73 | 204 | Glycosyl hydrolases family 39. |
| COG3664 | XynB | 2.38e-05 | 78 | 264 | 37 | 215 | Beta-xylosidase [Carbohydrate transport and metabolism]. |
| cd21510 | agarase_cat | 0.003 | 136 | 270 | 87 | 190 | alpha-beta barrel catalytic domain of agarase, such as GH86-like endo-acting agarases identified in non-marine organisms. Typically, agarases (E.C. 3.2.1.81) are found in ocean-dwelling bacteria since agarose is a principle component of red algae cell wall polysaccharides. Agarose is a linear polymer of alternating D-galactose and 3,6-anhydro-L-galactopyranose. Endo-acting agarases, such as glycoside hydrolase 16 (GH16) and GH86 hydrolyze internal beta-1,4 linkages. GH86-like endo-acting agarase of this protein family has been identified in the human intestinal bacterium Bacteroides uniformis. This acquired metabolic pathway, as demonstrated by the prevalence of agar-specific genetic cluster called polysaccharide utilization loci (PULs), varies considerably between human populations, being much more prevalent in a Japanese sample than in North America, European, or Chinese samples. Agarase activity was also identified in the non-marine bacterium Cellvibrio sp. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM69820.1 | 3.69e-115 | 1 | 402 | 1 | 388 |
| QJD82796.1 | 2.91e-70 | 1 | 401 | 1 | 378 |
| QJD85864.1 | 5.81e-67 | 2 | 403 | 3 | 386 |
| QTH43246.1 | 1.07e-65 | 1 | 401 | 1 | 378 |
| QEL16017.1 | 8.08e-53 | 14 | 365 | 38 | 368 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5Z3K_A | 5.66e-23 | 3 | 350 | 9 | 317 | Crystalstructure of glucosidase from Croceicoccus marinus at 1.8 Angstrom resolution [Croceicoccus marinus],5Z3K_B Crystal structure of glucosidase from Croceicoccus marinus at 1.8 Angstrom resolution [Croceicoccus marinus] |
| 1PX8_A | 9.63e-13 | 54 | 203 | 53 | 202 | Crystalstructure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1PX8_B Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_A Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_B Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_C Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_D Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O30360 | 5.27e-12 | 54 | 203 | 53 | 202 | Beta-xylosidase OS=Thermoanaerobacterium saccharolyticum (strain DSM 8691 / JW/SL-YS485) OX=1094508 GN=xynB PE=3 SV=1 |
| P36906 | 5.27e-12 | 54 | 203 | 53 | 202 | Beta-xylosidase OS=Thermoanaerobacterium saccharolyticum OX=28896 GN=xynB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
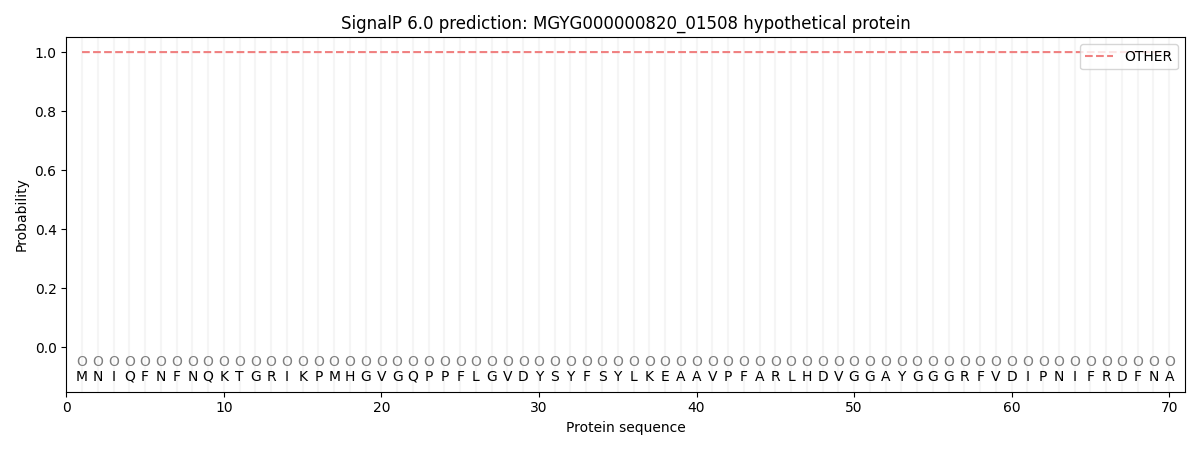
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000048 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
