You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000842_00015
Basic Information
help
| Species |
Collinsella sp900540845
|
| Lineage |
Bacteria; Actinobacteriota; Coriobacteriia; Coriobacteriales; Coriobacteriaceae; Collinsella; Collinsella sp900540845
|
| CAZyme ID |
MGYG000000842_00015
|
| CAZy Family |
GH140 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000000842 |
1846287 |
MAG |
China |
Asia |
|
| Gene Location |
Start: 17871;
End: 19175
Strand: +
|
No EC number prediction in MGYG000000842_00015.
| Family |
Start |
End |
Evalue |
family coverage |
| GH140 |
4 |
382 |
2.8e-85 |
0.8883495145631068 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| pfam13204
|
DUF4038 |
2.52e-86 |
18 |
336 |
9 |
317 |
Protein of unknown function (DUF4038). A family of putative cellulases. |
This protein is predicted as OTHER
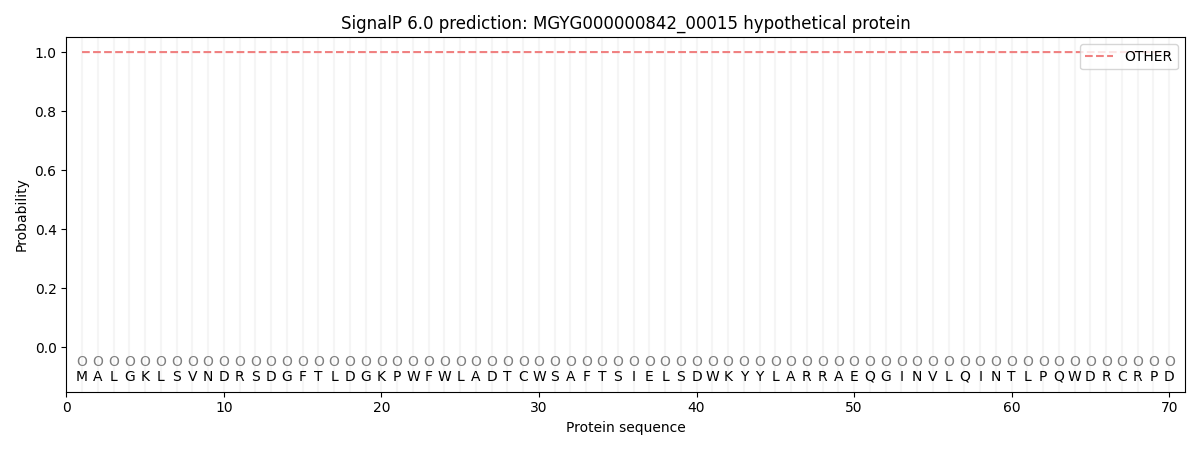
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000069
|
0.000002
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000000842_00015.

