You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000844_01122
Basic Information
help
| Species |
Lactobacillus apis
|
| Lineage |
Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Lactobacillus; Lactobacillus apis
|
| CAZyme ID |
MGYG000000844_01122
|
| CAZy Family |
GH38 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
| Protein Length |
CGC |
Molecular Weight |
Isoelectric Point |
| 181 |
|
20400.25 |
4.5359 |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000000844 |
1387413 |
MAG |
China |
Asia |
|
| Gene Location |
Start: 1944;
End: 2489
Strand: -
|
No EC number prediction in MGYG000000844_01122.
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| PRK09819
|
PRK09819 |
2.21e-10 |
3 |
138 |
706 |
822 |
mannosylglycerate hydrolase. |
| COG0383
|
AMS1 |
3.97e-04 |
25 |
172 |
789 |
932 |
Alpha-mannosidase [Carbohydrate transport and metabolism]. |
This protein is predicted as OTHER
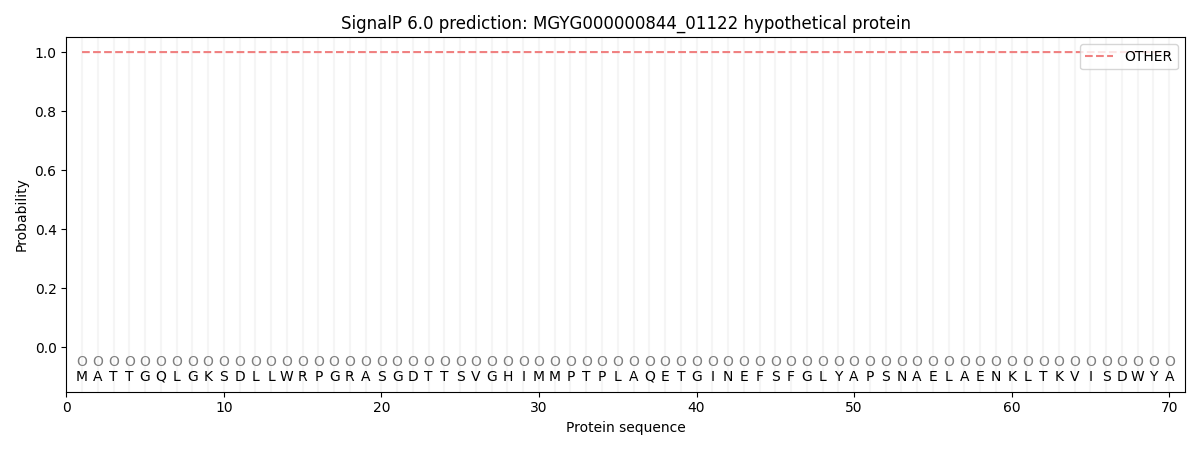
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000049
|
0.000001
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000000844_01122.

