You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000852_00812
You are here: Home > Sequence: MGYG000000852_00812
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900545525 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900545525 | |||||||||||
| CAZyme ID | MGYG000000852_00812 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 46412; End: 49285 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 478 | 639 | 3.9e-34 | 0.801980198019802 |
| CBM77 | 854 | 952 | 1.6e-23 | 0.941747572815534 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3866 | PelB | 5.36e-35 | 479 | 772 | 102 | 345 | Pectate lyase [Carbohydrate transport and metabolism]. |
| pfam18283 | CBM77 | 8.26e-29 | 854 | 954 | 7 | 108 | Carbohydrate binding module 77. This domain is the non-catalytic carbohydrate binding module 77 (CBM77) present in Ruminococcus flavefaciens. CBMs fulfil a critical targeting function in plant cell wall depolymerisation. In CBM77, a cluster of conserved basic residues (Lys1092, Lys1107 and Lys1162) confer calcium-independent recognition of homogalacturonan. |
| smart00656 | Amb_all | 2.87e-22 | 479 | 638 | 17 | 186 | Amb_all domain. |
| pfam00544 | Pec_lyase_C | 2.39e-08 | 474 | 638 | 30 | 211 | Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNT67442.1 | 5.27e-188 | 202 | 954 | 29 | 771 |
| QCD40816.1 | 3.45e-186 | 223 | 810 | 10 | 588 |
| QCP73706.1 | 3.45e-186 | 223 | 810 | 10 | 588 |
| QUT73893.1 | 1.35e-180 | 284 | 954 | 25 | 679 |
| QOR20273.1 | 6.40e-176 | 267 | 802 | 35 | 551 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5FU5_A | 9.39e-12 | 863 | 952 | 22 | 109 | Thecomplexity of the Ruminococcus flavefaciens cellulosome reflects an expansion in glycan recognition [Ruminococcus flavefaciens] |
| 1PCL_A | 1.31e-09 | 479 | 638 | 81 | 276 | ChainA, PECTATE LYASE E [Dickeya chrysanthemi] |
| 1VBL_A | 5.30e-09 | 479 | 638 | 133 | 330 | Structureof the thermostable pectate lyase PL 47 [Bacillus sp. TS-47] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q5AVN4 | 6.79e-15 | 479 | 651 | 99 | 277 | Pectate lyase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyA PE=1 SV=1 |
| B0XT32 | 2.15e-13 | 485 | 694 | 97 | 300 | Probable pectate lyase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyA PE=3 SV=1 |
| Q4WIT0 | 2.15e-13 | 485 | 694 | 97 | 300 | Probable pectate lyase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyA PE=3 SV=1 |
| A1CYB8 | 1.67e-12 | 485 | 694 | 97 | 300 | Probable pectate lyase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyA PE=3 SV=1 |
| A2QV36 | 2.36e-11 | 479 | 668 | 95 | 295 | Probable pectate lyase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=plyA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
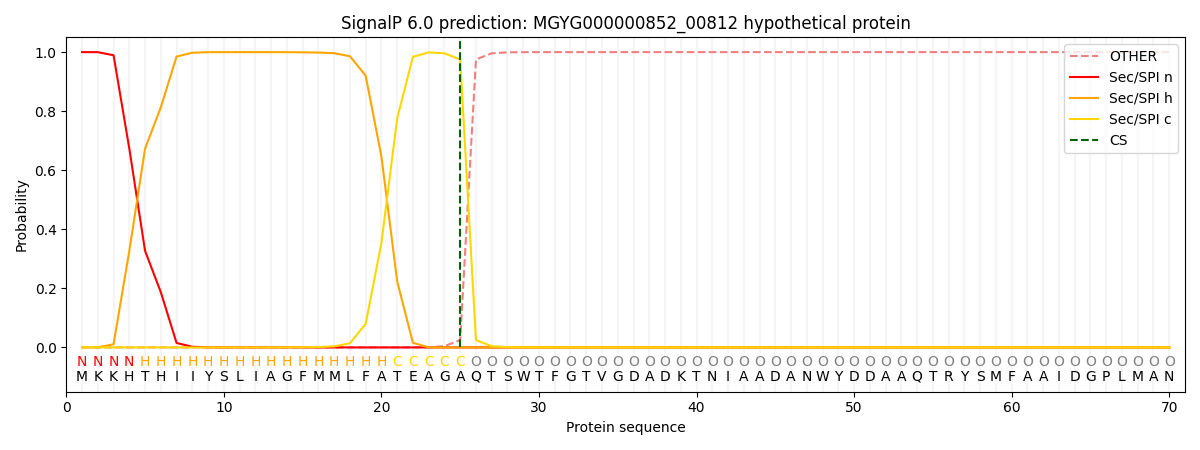
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000391 | 0.998709 | 0.000224 | 0.000239 | 0.000212 | 0.000195 |