You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000853_00500
You are here: Home > Sequence: MGYG000000853_00500
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900549175 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900549175 | |||||||||||
| CAZyme ID | MGYG000000853_00500 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 30276; End: 32519 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 473 | 734 | 1.7e-47 | 0.6798679867986799 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 2.30e-35 | 474 | 733 | 54 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 7.69e-34 | 471 | 735 | 93 | 310 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 2.84e-22 | 481 | 741 | 126 | 345 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
| pfam00331 | Glyco_hydro_10 | 1.41e-08 | 82 | 160 | 35 | 109 | Glycosyl hydrolase family 10. |
| smart00633 | Glyco_10 | 1.19e-06 | 91 | 141 | 1 | 50 | Glycosyl hydrolase family 10. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRX63290.1 | 9.50e-224 | 14 | 737 | 19 | 729 |
| ANU56191.1 | 6.99e-220 | 9 | 741 | 11 | 747 |
| QQR18969.1 | 6.99e-220 | 9 | 741 | 11 | 747 |
| AXH21505.1 | 2.34e-206 | 10 | 737 | 8 | 743 |
| EDV05072.1 | 2.34e-206 | 10 | 737 | 8 | 743 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1W2V_A | 1.14e-20 | 495 | 737 | 118 | 345 | The3-dimensional structure of a thermostable mutant of a xylanase (Xyn10A) from Cellvibrio japonicus [Cellvibrio japonicus],1W2V_B The 3-dimensional structure of a thermostable mutant of a xylanase (Xyn10A) from Cellvibrio japonicus [Cellvibrio japonicus] |
| 1CLX_A | 2.03e-20 | 495 | 737 | 117 | 344 | CatalyticCore Of Xylanase A [Cellvibrio japonicus],1CLX_B Catalytic Core Of Xylanase A [Cellvibrio japonicus],1CLX_C Catalytic Core Of Xylanase A [Cellvibrio japonicus],1CLX_D Catalytic Core Of Xylanase A [Cellvibrio japonicus] |
| 1W2P_A | 2.06e-20 | 495 | 737 | 118 | 345 | The3-dimensional structure of a xylanase (Xyn10A) from Cellvibrio japonicus [Cellvibrio japonicus],1W2P_B The 3-dimensional structure of a xylanase (Xyn10A) from Cellvibrio japonicus [Cellvibrio japonicus] |
| 1W32_A | 2.06e-20 | 495 | 737 | 118 | 345 | The3-dimensional structure of a thermostable mutant of a xylanase (Xyn10A) from Cellvibrio japonicus [Cellvibrio japonicus],1W32_B The 3-dimensional structure of a thermostable mutant of a xylanase (Xyn10A) from Cellvibrio japonicus [Cellvibrio japonicus] |
| 1W3H_A | 2.39e-20 | 495 | 737 | 129 | 356 | The3-dimensional structure of a thermostable mutant of a xylanase (Xyn10A) from Cellvibrio japonicus [Cellvibrio japonicus],1W3H_B The 3-dimensional structure of a thermostable mutant of a xylanase (Xyn10A) from Cellvibrio japonicus [Cellvibrio japonicus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P14768 | 7.36e-19 | 495 | 737 | 381 | 608 | Endo-1,4-beta-xylanase A OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=xynA PE=1 SV=2 |
| O69230 | 1.83e-16 | 475 | 679 | 472 | 656 | Endo-1,4-beta-xylanase C OS=Paenibacillus barcinonensis OX=198119 GN=xynC PE=1 SV=1 |
| Q9P8J1 | 5.21e-14 | 475 | 637 | 129 | 268 | Endo-1,4-beta-xylanase A OS=Talaromyces purpureogenus OX=1266744 GN=XynA PE=1 SV=1 |
| P56588 | 6.83e-14 | 475 | 637 | 102 | 241 | Endo-1,4-beta-xylanase OS=Penicillium simplicissimum OX=69488 PE=1 SV=1 |
| Q5S7A8 | 9.17e-14 | 475 | 637 | 127 | 266 | Endo-1,4-beta-xylanase A OS=Penicillium canescens OX=5083 GN=xylA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
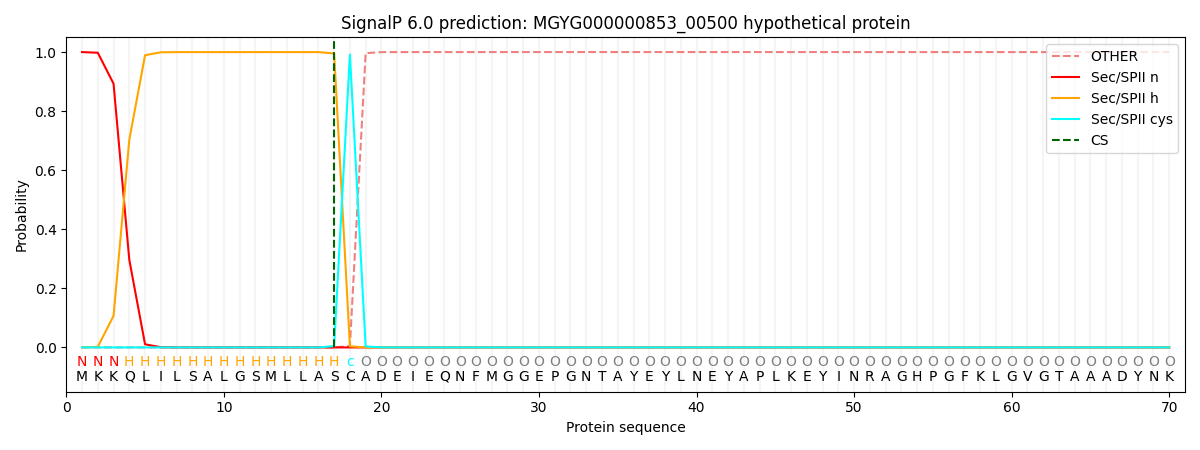
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000001 | 1.000058 | 0.000000 | 0.000000 | 0.000000 |
