You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000853_00596
You are here: Home > Sequence: MGYG000000853_00596
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900549175 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900549175 | |||||||||||
| CAZyme ID | MGYG000000853_00596 | |||||||||||
| CAZy Family | GH98 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7954; End: 10452 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH98 | 34 | 341 | 4e-77 | 0.9877675840978594 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam08306 | Glyco_hydro_98M | 5.26e-80 | 32 | 341 | 4 | 328 | Glycosyl hydrolase family 98. This domain is the putative catalytic domain of glycosyl hydrolase family 98 proteins. |
| cd04083 | CBM35_Lmo2446-like | 1.89e-12 | 647 | 778 | 2 | 125 | Carbohydrate Binding Module 35 (CBM35) domains similar to Lmo2446. This family includes carbohydrate binding module 35 (CBM35) domains that are appended to several carbohydrate binding enzymes. Some CBM35 domains belonging to this family are appended to glycoside hydrolase (GH) family domains, including glycoside hydrolase family 31 (GH31), for example the CBM35 domain of Lmo2446, an uncharacterized protein from Listeria monocytogenes EGD-e. These CBM35s are non-catalytic carbohydrate binding domains that facilitate the strong binding of the GH catalytic modules with their dedicated, insoluble substrates. GH31 has a wide range of hydrolytic activities such as alpha-glucosidase, alpha-xylosidase, 6-alpha-glucosyltransferase, or alpha-1,4-glucan lyase, cleaving a terminal carbohydrate moiety from a substrate that may be a starch or a glycoprotein. Most characterized GH31 enzymes are alpha-glucosidases. |
| pfam08307 | Glyco_hydro_98C | 2.78e-12 | 410 | 552 | 46 | 171 | Glycosyl hydrolase family 98 C-terminal domain. This putative domain is found at the C-terminus of glycosyl hydrolase family 98 proteins. This domain is not expected to form part of the catalytic activity. |
| cd04082 | CBM35_pectate_lyase-like | 5.05e-06 | 648 | 763 | 3 | 116 | Carbohydrate Binding Module family 35 (CBM35), pectate lyase-like; appended mainly to enzymes that bind mannan (Man), xylan, glucuronic acid (GlcA) and possibly glucans. This family includes carbohydrate binding module family 35 (CBM35) domains that are non-catalytic carbohydrate binding domains that are appended mainly to enzymes that bind mannan (Man), xylan, glucuronic acid (GlcA) and possibly glucans. Included in this family are CBM35s of pectate lyases, including pectate lyase 10A from Cellvibrio japonicas, these enzymes release delta-4,5-anhydrogalaturonic acid (delta4,5-GalA) from pectin, thus identifying a signature molecule for plant cell wall degradation. CBM35s are unique in that they display conserved specificity through extensive sequence similarity but divergent function through their appended catalytic modules. They are known to bind alpha-D-galactose (Gal), mannan (Man), xylan, glucuronic acid (GlcA), a beta-polymer of mannose, and possibly glucans, forming four subfamilies based on general ligand specificities (galacto, urono, manno, and gluco configurations). In contrast to most CBMs that are generally rigid proteins, CBM35 undergoes significant conformational change upon ligand binding. Some CBM35s bind their ligands in a calcium-dependent manner, especially those binding uronic acids. |
| cd04086 | CBM35_mannanase-like | 8.25e-05 | 668 | 758 | 17 | 107 | Carbohydrate Binding Module 35 (CBM35); appended to several carbohydrate binding enzymes, including several glycoside hydrolase (GH) family 26 mannanase domains. This family includes carbohydrate binding module 35 (CBM35) domains that are appended to several carbohydrate binding enzymes, including periplasmic component of ABC-type sugar transport system involved in carbohydrate transport and metabolism, and several glycoside hydrolase (GH) domains, including GH26. These CBM6s are non-catalytic carbohydrate binding domains that facilitate the strong binding of the GH catalytic modules with their dedicated, insoluble substrates. Examples of proteins having CMB35s belonging to this family are mannanase A from Clostridium thermocellum (GH26), Man26B from Paenibacillus sp. BME-14 (GH26), and the multifunctional Cel44C-Man26A from Paenibacillus polymyxa GS01 (which has two GH domains, GH44 and GH26). GH26 mainly includes mannan endo-1,4-beta-mannosidase which hydrolyzes 1,4-beta-D-linkages in mannans, galacto-mannans, glucomannans, and galactoglucomannans, but displays little activity towards other plant cell wall polysaccharides. A few proteins belonging to this family have additional CBM3 domains; these CBM3s are not found in the CBM6-CBM35-CBM36_like superfamily. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAV06735.1 | 3.98e-212 | 28 | 762 | 27 | 736 |
| AXY74108.1 | 3.75e-201 | 30 | 788 | 31 | 770 |
| ACE84677.1 | 1.87e-196 | 32 | 762 | 303 | 1004 |
| QEI20698.1 | 7.20e-196 | 32 | 762 | 351 | 1052 |
| QEI17121.1 | 7.20e-196 | 32 | 762 | 351 | 1052 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2WMF_A | 3.53e-09 | 29 | 552 | 34 | 488 | Crystalstructure of the catalytic module of a family 98 glycoside hydrolase from Streptococcus pneumoniae TIGR4 (Sp4GH98) in its native form. [Streptococcus pneumoniae TIGR4] |
| 2WMH_A | 3.53e-09 | 29 | 552 | 34 | 488 | Crystalstructure of the catalytic module of a family 98 glycoside hydrolase from Streptococcus pneumoniae TIGR4 in complex with the H- disaccharide blood group antigen. [Streptococcus pneumoniae TIGR4] |
| 2WMG_A | 8.07e-09 | 407 | 552 | 365 | 488 | Crystalstructure of the catalytic module of a family 98 glycoside hydrolase from Streptococcus pneumoniae TIGR4 (Sp4GH98) in complex with the LewisY pentasaccharide blood group antigen. [Streptococcus pneumoniae TIGR4] |
| 4D72_A | 6.03e-06 | 407 | 565 | 361 | 499 | Crystalstructure of a family 98 glycoside hydrolase catalytic module (Sp3GH98) in complex with the type 2 blood group A-tetrasaccharide (E558A L19 mutant) [Streptococcus pneumoniae SP3-BS71] |
| 4D71_A | 6.03e-06 | 407 | 565 | 361 | 499 | Crystalstructure of a family 98 glycoside hydrolase catalytic module (Sp3GH98) in complex with the type 2 blood group A-tetrasaccharide (E558A X02 mutant) [Streptococcus pneumoniae SP3-BS71] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
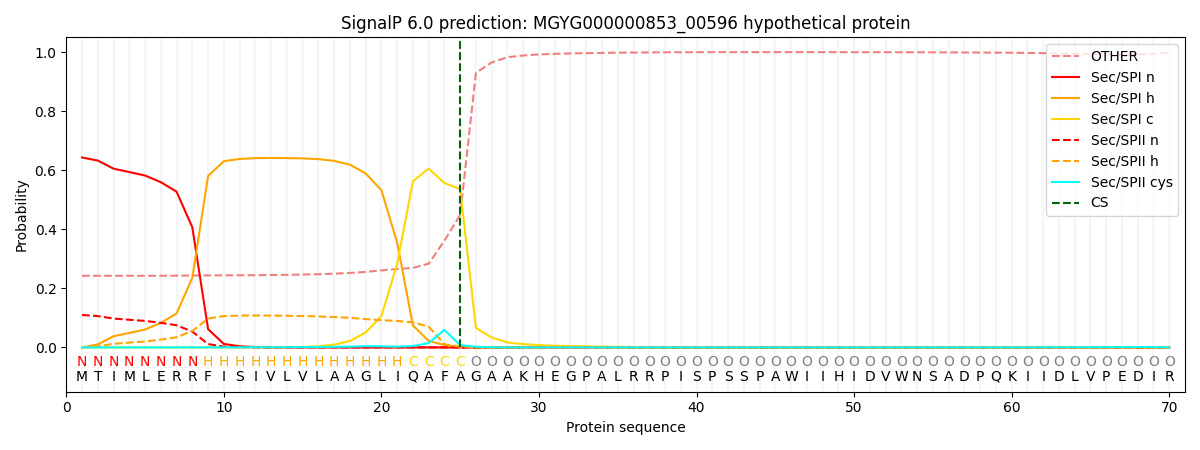
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.263935 | 0.612788 | 0.114346 | 0.006242 | 0.001464 | 0.001200 |