You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000853_01704
You are here: Home > Sequence: MGYG000000853_01704
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900549175 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900549175 | |||||||||||
| CAZyme ID | MGYG000000853_01704 | |||||||||||
| CAZy Family | PL4 | |||||||||||
| CAZyme Description | Rhamnogalacturonate lyase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 8430; End: 10319 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL4 | 22 | 565 | 9.1e-132 | 0.9963702359346642 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10317 | RGL4_C | 7.63e-41 | 414 | 568 | 1 | 161 | C-terminal domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold. Both the middle and the C-terminal domain are putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. |
| cd10320 | RGL4_N | 9.19e-40 | 24 | 281 | 3 | 262 | N-terminal catalytic domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold; the middle and C-terminal domains are both putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. |
| pfam14683 | CBM-like | 3.90e-34 | 413 | 567 | 2 | 157 | Polysaccharide lyase family 4, domain III. CBM-like is domain III of rhamnogalacturonan lyase (RG-lyase). The full-length protein specifically recognizes and cleaves alpha-1,4 glycosidic bonds between l-rhamnose and d-galacturonic acids in the backbone of rhamnogalacturonan-I, a major component of the plant cell wall polysaccharide, pectin. This domain possesses a jelly roll beta-sandwich fold structurally homologous to carbohydrate binding modules (CBMs), and it carries two sulfate ions and a hexa-coordinated calcium ion. |
| cd10316 | RGL4_M | 3.08e-18 | 311 | 401 | 1 | 91 | Middle domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold. Both the middle domain represented by this model and the C-terminal domain are putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. |
| pfam14686 | fn3_3 | 1.01e-09 | 333 | 398 | 10 | 74 | Polysaccharide lyase family 4, domain II. FnIII-like is domain II of rhamnogalacturonan lyase (RG-lyase). The full-length protein specifically recognizes and cleaves alpha-1,4 glycosidic bonds between l-rhamnose and d-galacturonic acids in the backbone of rhamnogalacturonan-I, a major component of the plant cell wall polysaccharide, pectin. This domain displays an immunoglobulin-like or more specifically Fibronectin-III type fold and shows highest structural similarity to the C-terminal beta-sandwich subdomain of the pro-hormone/propeptide processing enzyme carboxypeptidase gp180 from duck. It serves to assist in producing the deep pocket, with domain III, into which the substrate fits. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBH93641.1 | 1.23e-132 | 16 | 577 | 42 | 589 |
| BAV08848.1 | 4.56e-106 | 17 | 569 | 45 | 595 |
| APC39134.1 | 1.12e-104 | 20 | 569 | 3 | 563 |
| QOH51094.1 | 7.73e-97 | 2 | 567 | 9 | 571 |
| QOH46789.1 | 7.73e-97 | 2 | 567 | 9 | 571 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8RJP2 | 6.05e-93 | 2 | 567 | 11 | 573 | Rhamnogalacturonate lyase OS=Dickeya dadantii (strain 3937) OX=198628 GN=rhiE PE=1 SV=1 |
| Q5AZ85 | 6.28e-21 | 99 | 457 | 113 | 503 | Rhamnogalacturonate lyase B OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=rglB PE=2 SV=2 |
| Q0C7K7 | 8.32e-21 | 15 | 457 | 14 | 503 | Probable rhamnogalacturonate lyase B OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=rglB PE=3 SV=1 |
| B8N5T6 | 1.11e-20 | 99 | 491 | 114 | 558 | Probable rhamnogalacturonate lyase B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=rglB PE=3 SV=1 |
| A5ABH4 | 1.51e-19 | 105 | 548 | 120 | 632 | Probable rhamnogalacturonate lyase B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=rglB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
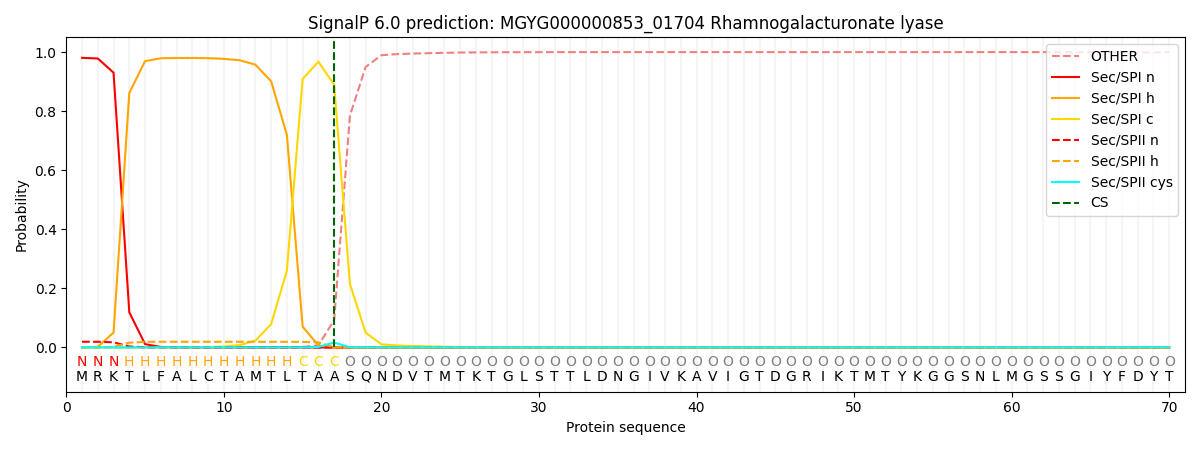
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001405 | 0.976852 | 0.020906 | 0.000287 | 0.000274 | 0.000263 |
