You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000867_01270
You are here: Home > Sequence: MGYG000000867_01270
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA1409 sp002338885 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; UBA1409; UBA1409 sp002338885 | |||||||||||
| CAZyme ID | MGYG000000867_01270 | |||||||||||
| CAZy Family | GH115 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 22278; End: 28025 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH115 | 70 | 758 | 3.1e-161 | 0.8020086083213773 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam15979 | Glyco_hydro_115 | 3.02e-120 | 236 | 630 | 3 | 334 | Glycosyl hydrolase family 115. Glyco_hydro_115 is a family of glycoside hydrolases likely to have the activity of xylan a-1,2-glucuronidase, EC:3.2.1.131, or a-(4-O-methyl)-glucuronidase EC:3.2.1.-. |
| cd01830 | XynE_like | 1.59e-43 | 1712 | 1907 | 1 | 200 | SGNH_hydrolase subfamily, similar to the putative arylesterase/acylhydrolase from the rumen anaerobe Prevotella bryantii XynE. The P. bryantii XynE gene is located in a xylanase gene cluster. SGNH hydrolases are a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| pfam17829 | GH115_C | 2.14e-26 | 895 | 1070 | 1 | 172 | Gylcosyl hydrolase family 115 C-terminal domain. This domain is found at the C-terminus of glycosyl hydrolase family 115 proteins. This domain has a beta-sandwich fold. |
| cd04501 | SGNH_hydrolase_like_4 | 5.87e-19 | 1713 | 1907 | 3 | 179 | Members of the SGNH-hydrolase superfamily, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad from other serine hydrolases, but may lack the carboxlic acid. |
| pfam13472 | Lipase_GDSL_2 | 6.53e-15 | 1715 | 1902 | 1 | 176 | GDSL-like Lipase/Acylhydrolase family. This family of presumed lipases and related enzymes are similar to pfam00657. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AUC17495.1 | 4.45e-163 | 46 | 1112 | 27 | 1050 |
| APZ46852.1 | 4.45e-163 | 46 | 1112 | 27 | 1050 |
| AFO77172.1 | 1.36e-146 | 74 | 1071 | 71 | 973 |
| ADJ45460.1 | 1.36e-146 | 74 | 1071 | 71 | 973 |
| AGT84300.1 | 1.36e-146 | 74 | 1071 | 71 | 973 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4ZMH_A | 1.91e-111 | 86 | 1072 | 24 | 934 | Crystalstructure of a five-domain GH115 alpha-Glucuronidase from the Marine Bacterium Saccharophagus degradans 2-40T [Saccharophagus degradans 2-40],4ZMH_B Crystal structure of a five-domain GH115 alpha-Glucuronidase from the Marine Bacterium Saccharophagus degradans 2-40T [Saccharophagus degradans 2-40] |
| 6NPS_A | 1.55e-93 | 130 | 1071 | 61 | 963 | Crystalstructure of GH115 enzyme AxyAgu115A from Amphibacillus xylanus [Amphibacillus xylanus NBRC 15112],6NPS_B Crystal structure of GH115 enzyme AxyAgu115A from Amphibacillus xylanus [Amphibacillus xylanus NBRC 15112] |
| 7PUG_A | 8.24e-76 | 131 | 750 | 66 | 625 | ChainA, xylan alpha-1,2-glucuronidase [uncultured bacterium] |
| 7PXQ_A | 2.11e-74 | 131 | 750 | 65 | 624 | ChainA, xylan alpha-1,2-glucuronidase [uncultured bacterium] |
| 4C90_A | 6.96e-73 | 74 | 756 | 65 | 646 | Evidencethat GH115 alpha-glucuronidase activity is dependent on conformational flexibility [Bacteroides ovatus],4C90_B Evidence that GH115 alpha-glucuronidase activity is dependent on conformational flexibility [Bacteroides ovatus],4C91_A Evidence that GH115 alpha-glucuronidase activity is dependent on conformational flexibility [Bacteroides ovatus],4C91_B Evidence that GH115 alpha-glucuronidase activity is dependent on conformational flexibility [Bacteroides ovatus] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as LIPO
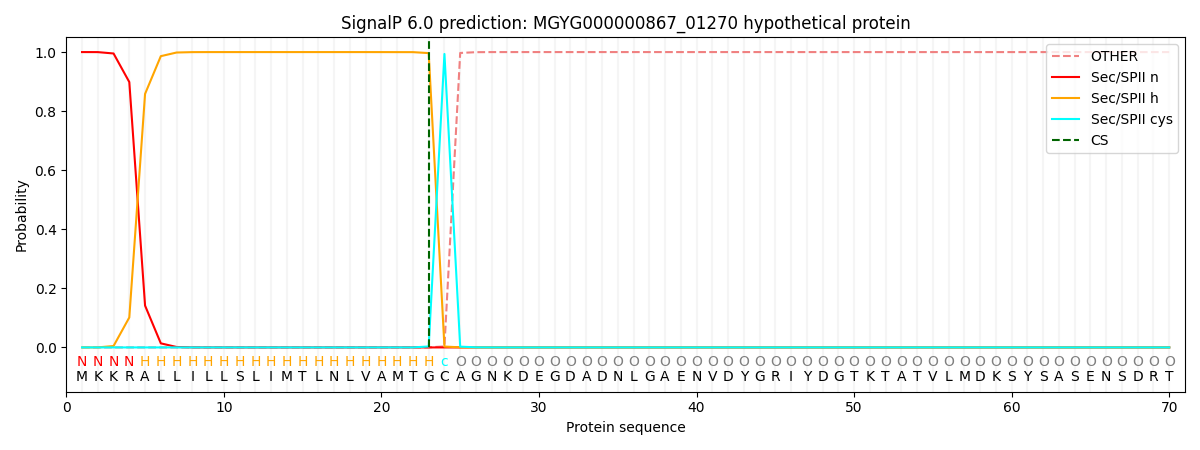
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000053 | 0.000000 | 0.000000 | 0.000000 |
