You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000922_00248
You are here: Home > Sequence: MGYG000000922_00248
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS416 sp900542005 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; UMGS416; UMGS416; UMGS416 sp900542005 | |||||||||||
| CAZyme ID | MGYG000000922_00248 | |||||||||||
| CAZy Family | GH109 | |||||||||||
| CAZyme Description | Alpha-N-acetylgalactosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 137508; End: 138632 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH109 | 2 | 365 | 6.9e-99 | 0.9774436090225563 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0673 | MviM | 3.35e-40 | 1 | 351 | 3 | 335 | Predicted dehydrogenase [General function prediction only]. |
| pfam01408 | GFO_IDH_MocA | 3.69e-17 | 2 | 117 | 1 | 119 | Oxidoreductase family, NAD-binding Rossmann fold. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
| PRK11579 | PRK11579 | 2.69e-07 | 23 | 145 | 29 | 149 | putative oxidoreductase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AGS53455.1 | 7.61e-179 | 1 | 372 | 1 | 372 |
| QHW35532.1 | 1.03e-171 | 2 | 372 | 3 | 381 |
| AEE96970.1 | 7.21e-168 | 1 | 370 | 1 | 371 |
| QHT62446.1 | 5.51e-167 | 2 | 372 | 3 | 382 |
| QHW33651.1 | 6.37e-166 | 2 | 372 | 3 | 382 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6T2B_A | 1.18e-38 | 8 | 360 | 51 | 433 | Glycosidehydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_B Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_C Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_D Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila] |
| 2IXA_A | 4.79e-36 | 3 | 371 | 20 | 437 | A-zyme,N-acetylgalactosaminidase [Elizabethkingia meningoseptica],2IXB_A Crystal structure of N-ACETYLGALACTOSAMINIDASE in complex with GalNAC [Elizabethkingia meningoseptica] |
| 4N54_A | 4.98e-11 | 2 | 342 | 15 | 336 | ChainA, Inositol dehydrogenase [Lacticaseibacillus casei BL23],4N54_B Chain B, Inositol dehydrogenase [Lacticaseibacillus casei BL23],4N54_C Chain C, Inositol dehydrogenase [Lacticaseibacillus casei BL23],4N54_D Chain D, Inositol dehydrogenase [Lacticaseibacillus casei BL23] |
| 4MKX_A | 5.05e-11 | 2 | 342 | 18 | 339 | ChainA, Inositol dehydrogenase [Lacticaseibacillus casei BL23],4MKZ_A Chain A, Inositol dehydrogenase [Lacticaseibacillus casei BL23] |
| 3MOI_A | 7.70e-09 | 25 | 141 | 29 | 145 | Thecrystal structure of the putative dehydrogenase from Bordetella bronchiseptica RB50 [Bordetella bronchiseptica] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A4Y8C8 | 4.79e-54 | 8 | 369 | 63 | 452 | Glycosyl hydrolase family 109 protein OS=Shewanella putrefaciens (strain CN-32 / ATCC BAA-453) OX=319224 GN=Sputcn32_2490 PE=3 SV=1 |
| A1RI61 | 6.71e-54 | 8 | 369 | 63 | 452 | Glycosyl hydrolase family 109 protein OS=Shewanella sp. (strain W3-18-1) OX=351745 GN=Sputw3181_1518 PE=3 SV=1 |
| A8H2K3 | 9.27e-53 | 8 | 369 | 61 | 449 | Glycosyl hydrolase family 109 protein OS=Shewanella pealeana (strain ATCC 700345 / ANG-SQ1) OX=398579 GN=Spea_1465 PE=3 SV=1 |
| A6WQ58 | 3.78e-52 | 8 | 369 | 63 | 452 | Glycosyl hydrolase family 109 protein OS=Shewanella baltica (strain OS185) OX=402882 GN=Shew185_2813 PE=3 SV=1 |
| A9KUT1 | 3.78e-52 | 8 | 369 | 63 | 452 | Glycosyl hydrolase family 109 protein OS=Shewanella baltica (strain OS195) OX=399599 GN=Sbal195_2933 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
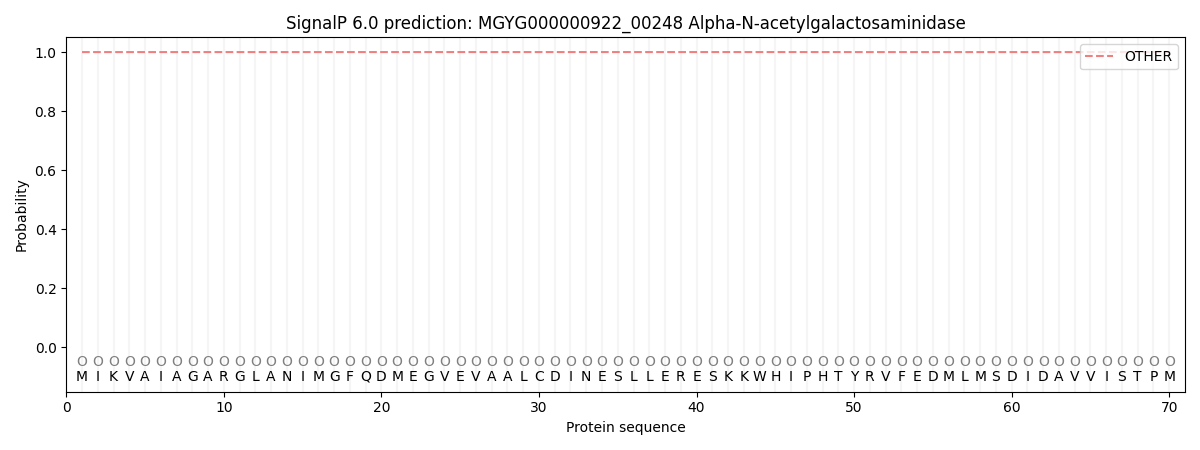
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000086 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
