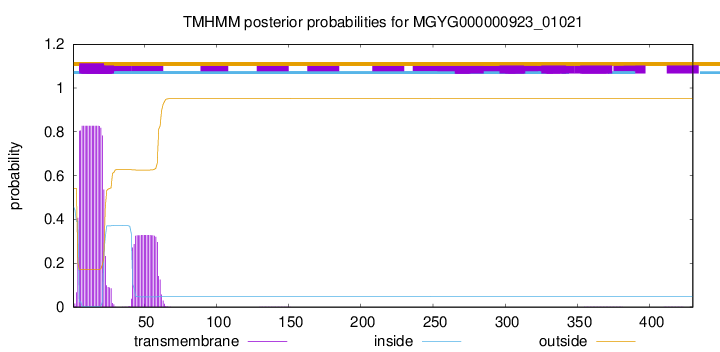You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000923_01021
You are here: Home > Sequence: MGYG000000923_01021
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola sp900552645 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola sp900552645 | |||||||||||
| CAZyme ID | MGYG000000923_01021 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 108225; End: 109517 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 65 | 377 | 5.3e-95 | 0.9930795847750865 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3934 | COG3934 | 1.38e-29 | 20 | 411 | 30 | 448 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
| COG3934 | COG3934 | 6.81e-15 | 35 | 333 | 4 | 247 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT89666.1 | 3.13e-206 | 15 | 421 | 11 | 421 |
| ALJ59276.1 | 8.95e-206 | 15 | 421 | 11 | 421 |
| QQA30547.1 | 2.60e-203 | 17 | 421 | 14 | 422 |
| ADD61406.1 | 2.60e-203 | 17 | 421 | 14 | 422 |
| ADD62011.1 | 2.60e-203 | 17 | 421 | 14 | 422 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1UUQ_A | 7.21e-100 | 20 | 421 | 16 | 423 | Exo-mannosidasefrom Cellvibrio mixtus [Cellvibrio mixtus],1UZ4_A Common inhibition of beta-glucosidase and beta-mannosidase by isofagomine lactam reflects different conformational intineraries for glucoside and mannoside hydrolysis [Cellvibrio mixtus],7ODJ_AAA Chain AAA, Man5A [Cellvibrio mixtus] |
| 4LYP_A | 6.70e-77 | 32 | 429 | 44 | 442 | CrystalStructure of Glycoside Hydrolase Family 5 Mannosidase from Rhizomucor miehei [Rhizomucor miehei],4LYP_B Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase from Rhizomucor miehei [Rhizomucor miehei] |
| 4LYQ_A | 5.21e-76 | 32 | 429 | 44 | 442 | CrystalStructure of Glycoside Hydrolase Family 5 Mannosidase from Rhizomucor miehei, E202A mutant [Rhizomucor miehei],4NRR_A Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase (E202A mutant) from Rhizomucor miehei in complex with mannosyl-fructose [Rhizomucor miehei],4NRR_B Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase (E202A mutant) from Rhizomucor miehei in complex with mannosyl-fructose [Rhizomucor miehei],4NRS_A Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase (E202A mutant) from Rhizomucor miehei in complex with mannobiose [Rhizomucor miehei],4NRS_B Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase (E202A mutant) from Rhizomucor miehei in complex with mannobiose [Rhizomucor miehei] |
| 4LYR_A | 5.21e-76 | 32 | 429 | 44 | 442 | GlycosideHydrolase Family 5 Mannosidase from Rhizomucor miehei, E301A mutant [Rhizomucor miehei] |
| 1RH9_A | 2.07e-33 | 102 | 331 | 72 | 294 | ChainA, endo-beta-mannanase [Solanum lycopersicum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9FJZ3 | 3.30e-34 | 23 | 391 | 27 | 390 | Mannan endo-1,4-beta-mannosidase 7 OS=Arabidopsis thaliana OX=3702 GN=MAN7 PE=2 SV=1 |
| O48540 | 6.55e-33 | 27 | 331 | 27 | 318 | Mannan endo-1,4-beta-mannosidase 1 OS=Solanum lycopersicum OX=4081 GN=MAN1 PE=1 SV=2 |
| Q8L5J1 | 1.78e-32 | 102 | 331 | 98 | 320 | Mannan endo-1,4-beta-mannosidase 4 OS=Solanum lycopersicum OX=4081 GN=MAN4 PE=1 SV=2 |
| Q9FZ29 | 2.79e-31 | 24 | 375 | 26 | 361 | Mannan endo-1,4-beta-mannosidase 1 OS=Arabidopsis thaliana OX=3702 GN=MAN1 PE=2 SV=1 |
| Q6YM50 | 4.01e-31 | 25 | 358 | 37 | 357 | Mannan endo-1,4-beta-mannosidase 5 OS=Solanum lycopersicum OX=4081 GN=MAN5 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000265 | 0.999105 | 0.000168 | 0.000155 | 0.000152 | 0.000137 |