You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000923_01335
You are here: Home > Sequence: MGYG000000923_01335
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola sp900552645 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola sp900552645 | |||||||||||
| CAZyme ID | MGYG000000923_01335 | |||||||||||
| CAZy Family | CBM67 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 32378; End: 34600 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH78 | 275 | 706 | 1.8e-78 | 0.876984126984127 |
| CBM67 | 34 | 196 | 2.6e-20 | 0.8920454545454546 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam17389 | Bac_rhamnosid6H | 2.43e-29 | 319 | 645 | 1 | 340 | Bacterial alpha-L-rhamnosidase 6 hairpin glycosidase domain. This family consists of bacterial rhamnosidase A and B enzymes. L-Rhamnose is abundant in biomass as a common constituent of glycolipids and glycosides, such as plant pigments, pectic polysaccharides, gums or biosurfactants. Some rhamnosides are important bioactive compounds. For example, terpenyl glycosides, the glycosidic precursor of aromatic terpenoids, act as important flavouring substances in grapes. Other rhamnosides act as cytotoxic rhamnosylated terpenoids, as signal substances in plants or play a role in the antigenicity of pathogenic bacteria. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNL36670.1 | 0.0 | 11 | 738 | 8 | 738 |
| QDH54696.1 | 0.0 | 7 | 738 | 6 | 738 |
| QLG44645.1 | 1.37e-254 | 31 | 723 | 31 | 731 |
| AGY53286.1 | 1.05e-244 | 28 | 726 | 31 | 739 |
| ACU03186.1 | 2.17e-243 | 18 | 730 | 28 | 749 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3CIH_A | 9.42e-23 | 296 | 685 | 276 | 684 | Crystalstructure of a putative alpha-rhamnosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
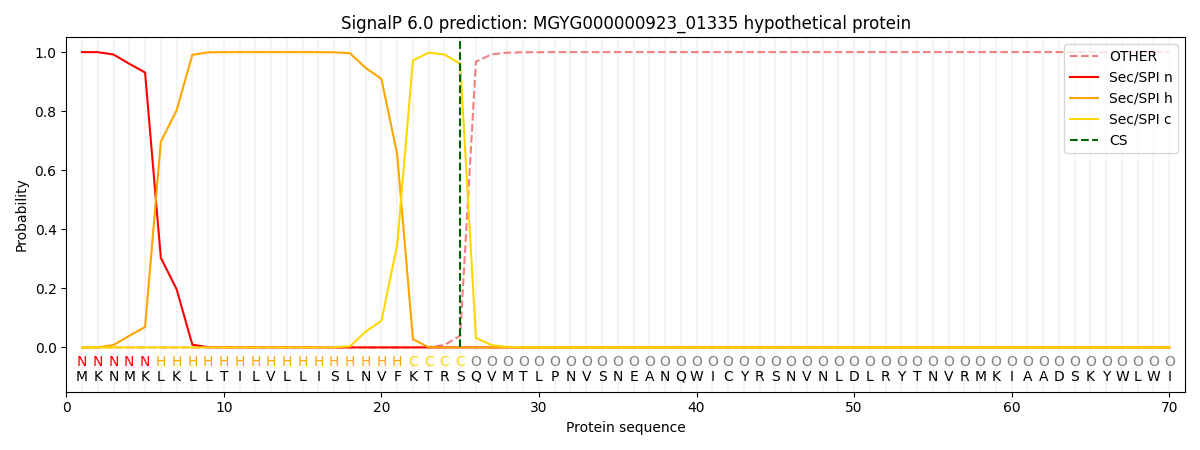
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000546 | 0.998624 | 0.000255 | 0.000219 | 0.000173 | 0.000164 |
