You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000964_01729
You are here: Home > Sequence: MGYG000000964_01729
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola salanitronis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola salanitronis | |||||||||||
| CAZyme ID | MGYG000000964_01729 | |||||||||||
| CAZy Family | CBM67 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7963; End: 11784 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH78 | 360 | 889 | 9.6e-162 | 0.9900793650793651 |
| CBM67 | 157 | 339 | 4e-40 | 0.9715909090909091 |
| GH33 | 923 | 1253 | 1.7e-36 | 0.9093567251461988 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam17389 | Bac_rhamnosid6H | 7.65e-127 | 464 | 821 | 3 | 339 | Bacterial alpha-L-rhamnosidase 6 hairpin glycosidase domain. This family consists of bacterial rhamnosidase A and B enzymes. L-Rhamnose is abundant in biomass as a common constituent of glycolipids and glycosides, such as plant pigments, pectic polysaccharides, gums or biosurfactants. Some rhamnosides are important bioactive compounds. For example, terpenyl glycosides, the glycosidic precursor of aromatic terpenoids, act as important flavouring substances in grapes. Other rhamnosides act as cytotoxic rhamnosylated terpenoids, as signal substances in plants or play a role in the antigenicity of pathogenic bacteria. |
| pfam13088 | BNR_2 | 3.82e-95 | 941 | 1251 | 1 | 280 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
| pfam08531 | Bac_rhamnosid_N | 5.97e-71 | 179 | 350 | 1 | 172 | Alpha-L-rhamnosidase N-terminal domain. This family consists of bacterial rhamnosidase A and B enzymes. This domain is probably involved in substrate recognition. |
| cd15482 | Sialidase_non-viral | 1.97e-51 | 920 | 1266 | 4 | 339 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| pfam05592 | Bac_rhamnosid | 8.07e-40 | 357 | 458 | 1 | 102 | Bacterial alpha-L-rhamnosidase concanavalin-like domain. This family consists of bacterial rhamnosidase A and B enzymes. L-Rhamnose is abundant in biomass as a common constituent of glycolipids and glycosides, such as plant pigments, pectic polysaccharides, gums or biosurfactants. Some rhamnosides are important bioactive compounds. For example, terpenyl glycosides, the glycosidic precursor of aromatic terpenoids, act as important flavouring substances in grapes. Other rhamnosides act as cytotoxic rhamnosylated terpenoids, as signal substances in plants or play a role in the antigenicity of pathogenic bacteria. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADY36775.1 | 0.0 | 1 | 1273 | 1 | 1273 |
| QIU95828.1 | 0.0 | 23 | 1266 | 18 | 1284 |
| QUR43127.1 | 0.0 | 11 | 1269 | 4 | 1285 |
| QGT69954.1 | 0.0 | 11 | 1269 | 4 | 1285 |
| QRQ56894.1 | 0.0 | 11 | 1269 | 4 | 1285 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6I60_A | 1.54e-178 | 19 | 890 | 14 | 899 | Structureof alpha-L-rhamnosidase from Dictyoglumus thermophilum [Dictyoglomus thermophilum H-6-12],6I60_B Structure of alpha-L-rhamnosidase from Dictyoglumus thermophilum [Dictyoglomus thermophilum H-6-12] |
| 3W5M_A | 7.16e-123 | 31 | 910 | 6 | 1034 | CrystalStructure of Streptomyces avermitilis alpha-L-rhamnosidase [Streptomyces avermitilis MA-4680 = NBRC 14893],3W5N_A Crystal Structure of Streptomyces avermitilis alpha-L-rhamnosidase complexed with L-rhamnose [Streptomyces avermitilis MA-4680 = NBRC 14893] |
| 6GSZ_A | 7.99e-101 | 29 | 904 | 2 | 863 | Crystalstructure of native alfa-L-rhamnosidase from Aspergillus terreus [Aspergillus terreus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KNB2 | 5.42e-137 | 23 | 897 | 29 | 886 | Alpha-L-rhamnosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22090 PE=1 SV=2 |
| T2KPL4 | 3.96e-134 | 40 | 897 | 41 | 914 | Alpha-L-rhamnosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22170 PE=2 SV=1 |
| Q82PP4 | 4.11e-122 | 31 | 897 | 6 | 1002 | Alpha-L-rhamnosidase OS=Streptomyces avermitilis (strain ATCC 31267 / DSM 46492 / JCM 5070 / NBRC 14893 / NCIMB 12804 / NRRL 8165 / MA-4680) OX=227882 GN=SAVERM_828 PE=1 SV=1 |
| P9WF03 | 4.72e-122 | 37 | 886 | 38 | 875 | Alpha-L-rhamnosidase OS=Alteromonas sp. (strain LOR) OX=1537994 GN=LOR_34 PE=1 SV=1 |
| T2KM26 | 7.99e-08 | 32 | 137 | 470 | 577 | Bifunctional sulfatase/alpha-L-rhamnosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22250 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
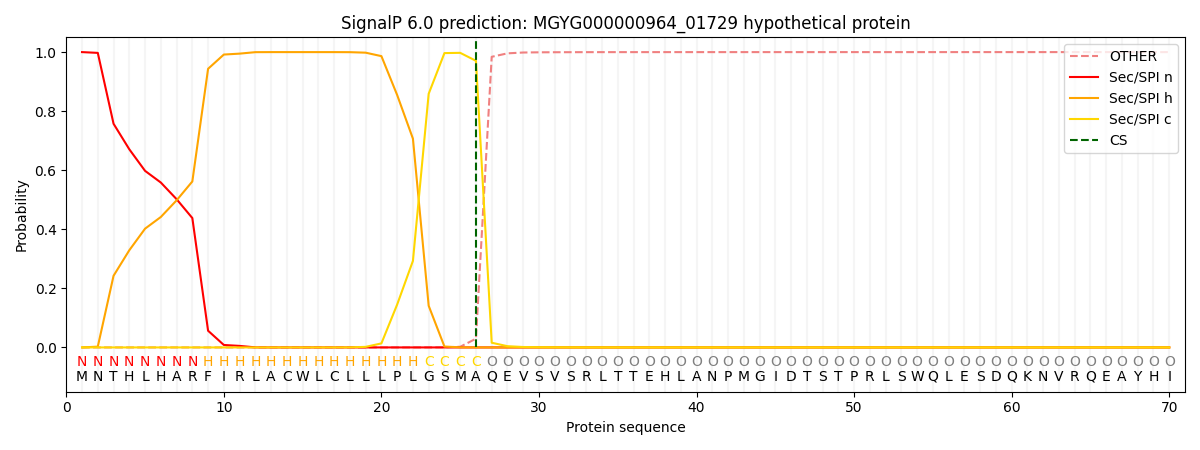
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000261 | 0.999062 | 0.000218 | 0.000166 | 0.000140 | 0.000134 |
