You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001000_00261
You are here: Home > Sequence: MGYG000001000_00261
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA4716 sp900556575 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales_A; UBA1381; UBA4716; UBA4716 sp900556575 | |||||||||||
| CAZyme ID | MGYG000001000_00261 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 185; End: 2962 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 28 | 555 | 1.9e-35 | 0.5478723404255319 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 2.58e-18 | 83 | 309 | 221 | 442 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10340 | ebgA | 1.11e-12 | 82 | 292 | 251 | 468 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| PRK09525 | lacZ | 4.61e-11 | 117 | 271 | 302 | 461 | beta-galactosidase. |
| pfam18565 | Glyco_hydro2_C5 | 5.05e-11 | 621 | 736 | 10 | 103 | Glycoside hydrolase family 2 C-terminal domain 5. Domain 5 is found in dimeric beta-D-galactosidase from Paracoccus sp. 32d, which contributes to stabilization of the functional dimer. It is suggested that the location of this domain 5, may be one of the factors responsible for the creation of a functional dimer and cold-adaptation of this enzyme. |
| pfam00703 | Glyco_hydro_2 | 1.42e-07 | 45 | 149 | 12 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ASA20342.1 | 1.39e-174 | 1 | 878 | 231 | 1045 |
| QGH35456.1 | 1.82e-170 | 1 | 903 | 238 | 1078 |
| AGT88372.1 | 1.23e-114 | 1 | 749 | 179 | 801 |
| ADJ49535.1 | 1.23e-114 | 1 | 749 | 179 | 801 |
| QKJ29765.1 | 2.36e-104 | 1 | 756 | 221 | 891 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6HPD_A | 1.94e-74 | 1 | 772 | 215 | 837 | Thestructure of a beta-glucuronidase from glycoside hydrolase family 2 [Formosa agariphila KMM 3901] |
| 5DMY_A | 9.37e-14 | 105 | 742 | 265 | 885 | Beta-galactosidase- construct 33-930 [Bifidobacterium bifidum],5DMY_B Beta-galactosidase - construct 33-930 [Bifidobacterium bifidum],5DMY_C Beta-galactosidase - construct 33-930 [Bifidobacterium bifidum] |
| 7CWD_A | 1.16e-13 | 20 | 741 | 184 | 803 | ChainA, beta-glalactosidase [Niallia circulans],7CWI_A Chain A, beta-galactosidase [Niallia circulans] |
| 6ED1_A | 4.75e-13 | 115 | 494 | 265 | 637 | ChainA, Glycosyl hydrolase family 2, sugar binding domain protein [Phocaeicola dorei],6ED1_B Chain B, Glycosyl hydrolase family 2, sugar binding domain protein [Phocaeicola dorei],6ED1_C Chain C, Glycosyl hydrolase family 2, sugar binding domain protein [Phocaeicola dorei],6ED1_D Chain D, Glycosyl hydrolase family 2, sugar binding domain protein [Phocaeicola dorei] |
| 6QUB_B | 7.21e-12 | 105 | 742 | 236 | 856 | Truncatedbeta-galactosidase III from Bifidobacterium bifidum in complex with galactose [Bifidobacterium bifidum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KN75 | 9.57e-74 | 1 | 772 | 206 | 828 | Beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22060 PE=1 SV=1 |
| A4W7D2 | 9.43e-13 | 105 | 306 | 291 | 497 | Beta-galactosidase OS=Enterobacter sp. (strain 638) OX=399742 GN=lacZ PE=3 SV=1 |
| A3FEW8 | 1.63e-12 | 105 | 306 | 291 | 497 | Beta-galactosidase OS=Enterobacter agglomerans OX=549 GN=lacZ PE=1 SV=2 |
| A5F5U6 | 1.10e-11 | 117 | 306 | 299 | 493 | Beta-galactosidase OS=Vibrio cholerae serotype O1 (strain ATCC 39541 / Classical Ogawa 395 / O395) OX=345073 GN=lacZ PE=3 SV=2 |
| Q2XQU3 | 1.67e-10 | 117 | 306 | 303 | 497 | Beta-galactosidase 2 OS=Enterobacter cloacae OX=550 GN=lacZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
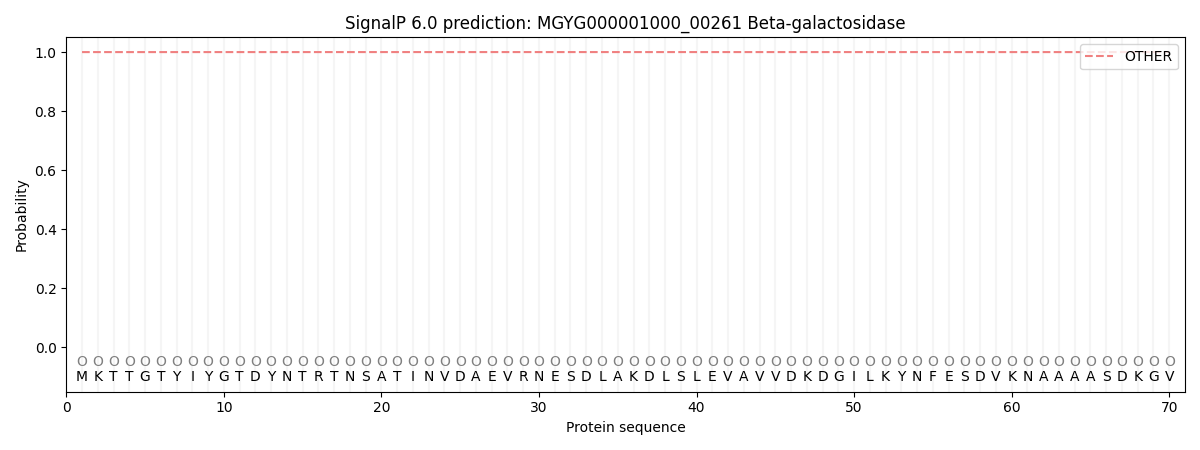
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000066 | 0.000002 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
