You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001014_01025
You are here: Home > Sequence: MGYG000001014_01025
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
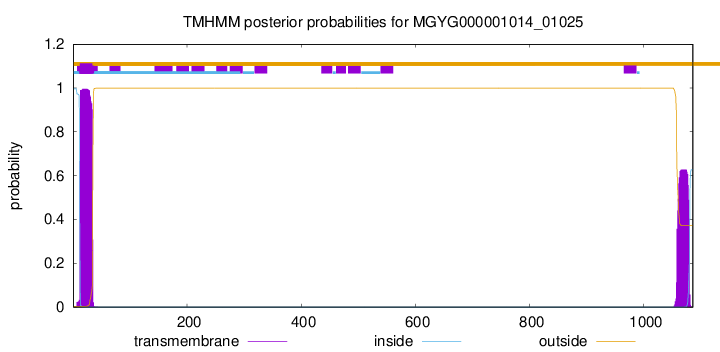
TMHMM annotations
Basic Information help
| Species | Varibaculum timonense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Actinomycetaceae; Varibaculum; Varibaculum timonense | |||||||||||
| CAZyme ID | MGYG000001014_01025 | |||||||||||
| CAZy Family | GH33 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1240; End: 4500 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH33 | 422 | 772 | 1.1e-72 | 0.9239766081871345 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd15482 | Sialidase_non-viral | 5.03e-74 | 430 | 783 | 15 | 339 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| COG4409 | NanH | 1.25e-22 | 378 | 746 | 221 | 613 | Neuraminidase (sialidase) [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| pfam13859 | BNR_3 | 1.08e-10 | 439 | 680 | 8 | 211 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
| pfam05345 | He_PIG | 1.44e-09 | 900 | 981 | 1 | 94 | Putative Ig domain. This alignment represents the conserved core region of ~90 residue repeat found in several haemagglutinins and other cell surface proteins. Sequence similarities to (pfam02494) and (pfam00801) suggest an Ig-like fold (personal obs:C. Yeats). So this family may be similar in function to the (pfam02639) and (pfam02638) domains. This domain is also found in the WisP family of proteins of Tropheryma whipplei. |
| NF033186 | internalin_K | 1.73e-08 | 856 | 1083 | 399 | 604 | class 1 internalin InlK. Internalins, as found in the intracellular human pathogen Listeria monocytogenes, are paralogous surface-anchored proteins with an N-terminal signal peptide, leucine-rich repeats, and a C-terminal LPXTG processing and cell surface anchoring site. Members of this family are internalin K (InlK), a virulence factor. See articles PMID:17764999. for a general discussion of internalins, and PMID:21829365, PMID:22082958, and PMID:23958637 for more information about internalin K. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AZR06434.1 | 9.31e-208 | 32 | 891 | 52 | 917 |
| AWG04339.1 | 2.17e-197 | 32 | 891 | 52 | 918 |
| AZR04057.1 | 2.17e-197 | 32 | 891 | 52 | 918 |
| AWG17066.1 | 2.17e-197 | 32 | 891 | 52 | 918 |
| AZR00725.1 | 8.31e-192 | 59 | 891 | 73 | 919 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1EUR_A | 5.92e-83 | 428 | 787 | 24 | 359 | Sialidase[Micromonospora viridifaciens],1EUS_A Sialidase Complexed With 2-Deoxy-2,3-Dehydro-N- Acetylneuraminic Acid [Micromonospora viridifaciens] |
| 7LBU_A | 3.23e-82 | 430 | 812 | 95 | 452 | ChainA, Exo-alpha-sialidase [Cutibacterium acnes],7LBV_A Chain A, Exo-alpha-sialidase [Cutibacterium acnes] |
| 1EUT_A | 4.33e-80 | 428 | 787 | 24 | 359 | Sialidase,Large 68kd Form, Complexed With Galactose [Micromonospora viridifaciens],1EUU_A Sialidase Or Neuraminidase, Large 68kd Form [Micromonospora viridifaciens] |
| 1WCQ_A | 1.41e-79 | 428 | 787 | 20 | 355 | Mutagenesisof the Nucleophilic Tyrosine in a Bacterial Sialidase to Phenylalanine. [Micromonospora viridifaciens],1WCQ_B Mutagenesis of the Nucleophilic Tyrosine in a Bacterial Sialidase to Phenylalanine. [Micromonospora viridifaciens],1WCQ_C Mutagenesis of the Nucleophilic Tyrosine in a Bacterial Sialidase to Phenylalanine. [Micromonospora viridifaciens] |
| 2BZD_A | 2.67e-79 | 428 | 787 | 20 | 355 | Galactoserecognition by the carbohydrate-binding module of a bacterial sialidase. [Micromonospora viridifaciens],2BZD_B Galactose recognition by the carbohydrate-binding module of a bacterial sialidase. [Micromonospora viridifaciens],2BZD_C Galactose recognition by the carbohydrate-binding module of a bacterial sialidase. [Micromonospora viridifaciens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02834 | 6.33e-79 | 428 | 787 | 66 | 401 | Sialidase OS=Micromonospora viridifaciens OX=1881 GN=nedA PE=1 SV=1 |
| P15698 | 9.35e-13 | 430 | 771 | 55 | 383 | Sialidase OS=Paeniclostridium sordellii OX=1505 PE=1 SV=1 |
| Q6GDE9 | 2.61e-11 | 805 | 977 | 587 | 744 | Serine-rich adhesin for platelets OS=Staphylococcus aureus (strain MRSA252) OX=282458 GN=sasA PE=3 SV=1 |
| Q8VQ99 | 2.91e-11 | 805 | 977 | 587 | 744 | Serine-rich adhesin for platelets OS=Staphylococcus aureus OX=1280 GN=sraP PE=3 SV=1 |
| Q8NUJ3 | 3.81e-11 | 805 | 977 | 587 | 744 | Serine-rich adhesin for platelets OS=Staphylococcus aureus (strain MW2) OX=196620 GN=sraP PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
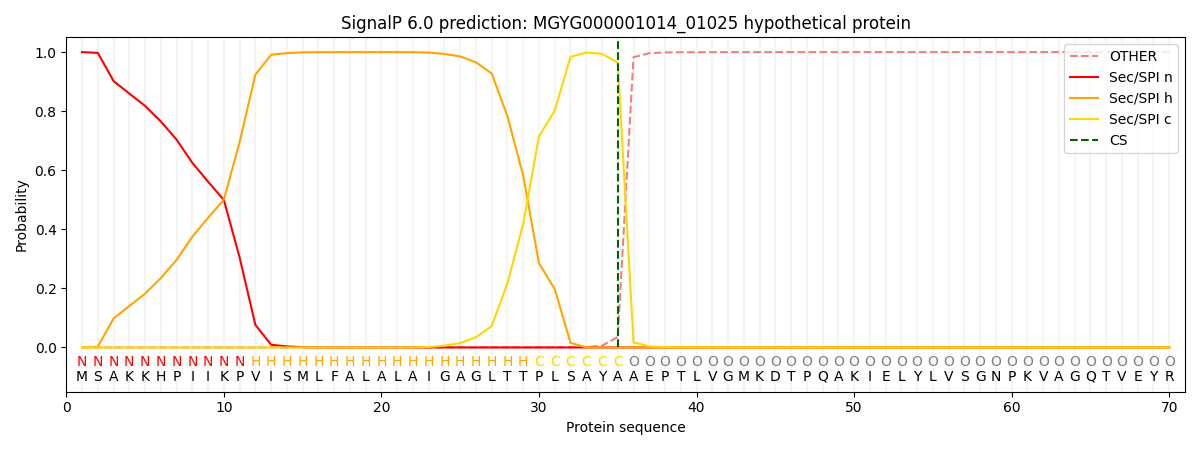
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000491 | 0.998552 | 0.000251 | 0.000289 | 0.000201 | 0.000173 |