You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001056_00652
You are here: Home > Sequence: MGYG000001056_00652
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900556395 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900556395 | |||||||||||
| CAZyme ID | MGYG000001056_00652 | |||||||||||
| CAZy Family | GH88 | |||||||||||
| CAZyme Description | Unsaturated chondroitin disaccharide hydrolase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 16827; End: 18029 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH88 | 70 | 394 | 3.6e-120 | 0.9878419452887538 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07470 | Glyco_hydro_88 | 2.94e-12 | 72 | 270 | 24 | 207 | Glycosyl Hydrolase Family 88. Unsaturated glucuronyl hydrolase catalyzes the hydrolytic release of unsaturated glucuronic acids from oligosaccharides (EC:3.2.1.-) produced by the reactions of polysaccharide lyases. |
| cd04434 | LanC_like | 0.002 | 66 | 180 | 35 | 160 | Cyclases involved in the biosynthesis of lantibiotics, and similar proteins. LanC is the cyclase enzyme of the lanthionine synthetase. Lanthionine is a lantibiotic, a unique class of peptide antibiotics. They are ribosomally synthesized as a precursor peptide and then post-translationally modified to contain thioether cross-links called lanthionines (Lans) or methyllanthionines (MeLans), in addition to 2,3-didehydroalanine (Dha) and (Z)-2,3-didehydrobutyrine (Dhb). These unusual amino acids are introduced by the dehydration of serine and threonine residues, followed by thioether formation via addition of cysteine thiols, catalysed by LanB and LanC or LanM. LanC, the cyclase component, is a zinc metalloprotein, whose bound metal has been proposed to activate the thiol substrate for nucleophilic addition. A related domain is also present in LanM and other pro- and eukaryotic proteins with poorly characterized functions. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AYD49376.1 | 1.51e-151 | 34 | 397 | 27 | 390 |
| AHF16959.1 | 5.01e-145 | 35 | 397 | 25 | 389 |
| AWK07512.1 | 2.68e-144 | 34 | 397 | 21 | 384 |
| QUR43442.1 | 1.26e-142 | 38 | 397 | 26 | 385 |
| QUT82307.1 | 1.76e-142 | 38 | 397 | 26 | 385 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3WIW_A | 6.15e-86 | 59 | 393 | 56 | 385 | Crystalstructure of unsaturated glucuronyl hydrolase specific for heparin [Pedobacter heparinus DSM 2366] |
| 1VD5_A | 5.52e-54 | 58 | 394 | 24 | 362 | CrystalStructure of Unsaturated Glucuronyl Hydrolase, Responsible for the Degradation of Glycosaminoglycan, from Bacillus sp. GL1 at 1.8 A Resolution [Bacillus sp. GL1],2D5J_A Unsaturated Glucuronyl Hydrolase Triggers Hydration of Vinyl Ether Group but not of Glycosidic Bond [Bacillus sp. GL1],2D5J_B Unsaturated Glucuronyl Hydrolase Triggers Hydration of Vinyl Ether Group but not of Glycosidic Bond [Bacillus sp. GL1],2FUZ_A UGL hexagonal crystal structure without glycine and DTT molecules [Bacillus sp. GL1] |
| 2AHF_A | 2.99e-53 | 58 | 394 | 24 | 362 | ChainA, unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2AHF_B Chain B, unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2AHG_A Chain A, unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2AHG_B Chain B, unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2FV0_A Chain A, Unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2FV0_B Chain B, Unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2FV1_A Chain A, Unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2FV1_B Chain B, Unsaturated glucuronyl hydrolase [Bacillus sp. GL1] |
| 2ZZR_A | 6.79e-53 | 66 | 394 | 57 | 387 | Crystalstructure of unsaturated glucuronyl hydrolase from Streptcoccus agalactiae [Streptococcus agalactiae] |
| 3ANJ_A | 6.95e-53 | 66 | 394 | 58 | 388 | Crystalstructure of unsaturated glucuronyl hydrolase from Streptcoccus agalactiae [Streptococcus agalactiae serogroup III] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KLZ3 | 8.94e-85 | 58 | 393 | 58 | 391 | Unsaturated glucuronyl hydrolase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21900 PE=1 SV=1 |
| Q8DR77 | 1.17e-54 | 33 | 394 | 37 | 386 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=ugl PE=1 SV=1 |
| Q9RC92 | 3.02e-53 | 58 | 394 | 24 | 362 | Unsaturated glucuronyl hydrolase OS=Bacillus sp. (strain GL1) OX=84635 GN=ugl PE=1 SV=1 |
| Q9A0T3 | 2.78e-52 | 27 | 394 | 22 | 389 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus pyogenes serotype M1 OX=301447 GN=ugl PE=1 SV=1 |
| Q8E372 | 3.81e-52 | 66 | 394 | 58 | 388 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus agalactiae serotype III (strain NEM316) OX=211110 GN=gbs1889 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
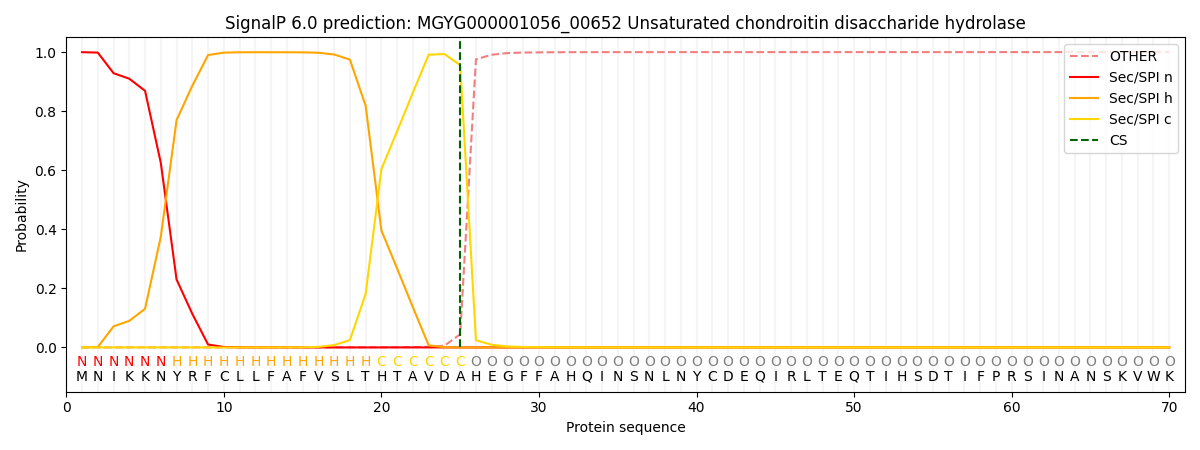
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000985 | 0.998070 | 0.000361 | 0.000201 | 0.000180 | 0.000181 |
