You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001063_03877
You are here: Home > Sequence: MGYG000001063_03877
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
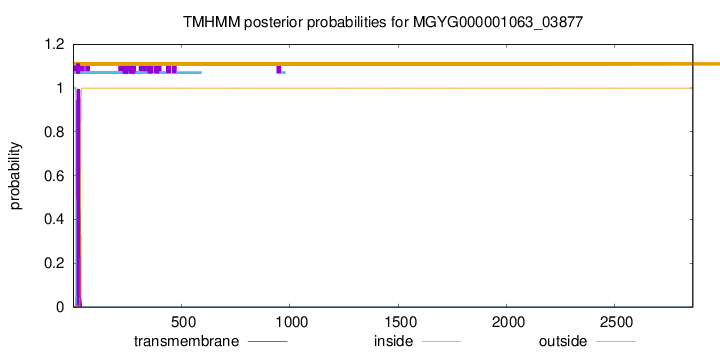
TMHMM annotations
Basic Information help
| Species | Robinsoniella sp900540475 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Robinsoniella; Robinsoniella sp900540475 | |||||||||||
| CAZyme ID | MGYG000001063_03877 | |||||||||||
| CAZy Family | CBM40 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4911; End: 13496 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH33 | 1333 | 1774 | 1.1e-90 | 0.9532163742690059 |
| CBM40 | 1152 | 1309 | 2.6e-35 | 0.9050279329608939 |
| CBM51 | 2105 | 2252 | 1.4e-20 | 0.9776119402985075 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd15482 | Sialidase_non-viral | 1.36e-76 | 1332 | 1777 | 1 | 339 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| COG4409 | NanH | 1.81e-28 | 1070 | 1771 | 18 | 706 | Neuraminidase (sialidase) [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| pfam08305 | NPCBM | 7.13e-23 | 2101 | 2252 | 1 | 135 | NPCBM/NEW2 domain. This novel putative carbohydrate binding module (NPCBM) domain is found at the N-terminus of glycosyl hydrolase family 98 proteins. This domain has also been called the NEW2 domain (Naumoff DG. Phylogenetic analysis of alpha-galactosidases of the GH27 family. Molecular Biology (Engl Transl). (2004)38:388-399.) |
| cd00229 | SGNH_hydrolase | 1.52e-21 | 704 | 907 | 1 | 186 | SGNH_hydrolase, or GDSL_hydrolase, is a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the typical Ser-His-Asp(Glu) triad from other serine hydrolases, but may lack the carboxlic acid. |
| pfam13472 | Lipase_GDSL_2 | 6.03e-21 | 706 | 898 | 1 | 174 | GDSL-like Lipase/Acylhydrolase family. This family of presumed lipases and related enzymes are similar to pfam00657. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ATD57534.1 | 2.34e-124 | 1155 | 1877 | 210 | 942 |
| QBJ75069.1 | 2.34e-124 | 1155 | 1877 | 210 | 942 |
| ATD54786.1 | 2.34e-124 | 1155 | 1877 | 210 | 942 |
| SLK16343.1 | 2.34e-124 | 1155 | 1877 | 210 | 942 |
| QJA08933.1 | 7.62e-122 | 1095 | 1848 | 13 | 752 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2BF6_A | 1.36e-96 | 1326 | 1783 | 4 | 448 | AtomicResolution Structure of the bacterial sialidase NanI from Clostridium perfringens in complex with alpha-Sialic Acid (Neu5Ac). [Clostridium perfringens] |
| 2VK5_A | 1.49e-96 | 1326 | 1783 | 4 | 448 | TheStructure Of Clostridium Perfringens Nani Sialidase And Its Catalytic Intermediates [Clostridium perfringens],2VK6_A The Structure Of Clostridium Perfringens Nani Sialidase And Its Catalytic Intermediates [Clostridium perfringens],2VK7_A The Structure Of Clostridium Perfringens Nani Sialidase And Its Catalytic Intermediates [Clostridium perfringens],2VK7_B The Structure Of Clostridium Perfringens Nani Sialidase And Its Catalytic Intermediates [Clostridium perfringens] |
| 5TSP_A | 3.63e-96 | 1326 | 1783 | 5 | 449 | Crystalstructure of the catalytic domain of Clostridium perfringens neuraminidase (NanI) in complex with a CHES [Clostridium perfringens ATCC 13124],5TSP_B Crystal structure of the catalytic domain of Clostridium perfringens neuraminidase (NanI) in complex with a CHES [Clostridium perfringens ATCC 13124] |
| 5KKY_A | 3.35e-83 | 1322 | 1784 | 21 | 496 | Structureof Streptococcus pneumonia NanA bound with inhibitor 9N3Neu5Ac2en [Streptococcus pneumoniae],5KKY_B Structure of Streptococcus pneumonia NanA bound with inhibitor 9N3Neu5Ac2en [Streptococcus pneumoniae] |
| 2YA4_A | 7.70e-83 | 1323 | 1784 | 19 | 493 | Crystalstructure of Streptococcus pneumoniae NanA (TIGR4) [Streptococcus pneumoniae TIGR4],2YA4_B Crystal structure of Streptococcus pneumoniae NanA (TIGR4) [Streptococcus pneumoniae TIGR4],2YA5_A Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with sialic acid [Streptococcus pneumoniae TIGR4],2YA5_B Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with sialic acid [Streptococcus pneumoniae TIGR4],2YA6_A Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with DANA [Streptococcus pneumoniae TIGR4],2YA6_B Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with DANA [Streptococcus pneumoniae TIGR4],2YA7_A Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with Zanamivir [Streptococcus pneumoniae TIGR4],2YA7_B Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with Zanamivir [Streptococcus pneumoniae TIGR4],2YA7_C Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with Zanamivir [Streptococcus pneumoniae TIGR4],2YA7_D Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with Zanamivir [Streptococcus pneumoniae TIGR4],2YA8_A Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with Oseltamivir carboxylate [Streptococcus pneumoniae TIGR4],2YA8_B Crystal structure of Streptococcus pneumoniae NanA (TIGR4) in complex with Oseltamivir carboxylate [Streptococcus pneumoniae TIGR4] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P29767 | 4.61e-115 | 1133 | 1842 | 190 | 891 | Sialidase OS=Clostridium septicum OX=1504 PE=3 SV=1 |
| P62576 | 5.23e-87 | 1141 | 1848 | 128 | 849 | Sialidase A OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=nanA PE=1 SV=1 |
| P62575 | 5.23e-87 | 1141 | 1848 | 128 | 849 | Sialidase A OS=Streptococcus pneumoniae OX=1313 GN=nanA PE=1 SV=1 |
| Q27701 | 7.79e-68 | 1116 | 1765 | 58 | 729 | Anhydrosialidase OS=Macrobdella decora OX=6405 PE=1 SV=1 |
| Q54727 | 5.04e-35 | 1159 | 1765 | 62 | 669 | Sialidase B OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=nanB PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000863 | 0.998113 | 0.000464 | 0.000174 | 0.000177 | 0.000166 |