You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001065_00929
You are here: Home > Sequence: MGYG000001065_00929
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Enterocloster sp900538485 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Enterocloster; Enterocloster sp900538485 | |||||||||||
| CAZyme ID | MGYG000001065_00929 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 77076; End: 78248 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 120 | 360 | 8.2e-27 | 0.9580152671755725 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 4.43e-14 | 123 | 350 | 27 | 258 | Cellulase (glycosyl hydrolase family 5). |
| COG5263 | COG5263 | 9.86e-09 | 26 | 83 | 257 | 313 | Glucan-binding domain (YG repeat) [Carbohydrate transport and metabolism]. |
| pfam19085 | Choline_bind_2 | 9.08e-06 | 43 | 82 | 3 | 38 | Choline-binding repeat. this entry contains a pair of presumed choline-binding repeats that are often found adjacent to pfam01473. |
| NF033840 | PspC_relate_1 | 1.21e-05 | 26 | 83 | 592 | 647 | PspC-related protein choline-binding protein 1. Members of this family share C-terminal homology to the choline-binding form of the pneumococcal surface antigen PspC, but not to its allelic LPXTG-anchored forms because they lack the choline-binding repeat region. Members of this family should not be confused with PspC itself, whose identity and function reflect regions N-terminal to the choline-binding region. See Iannelli, et al. (PMID: 11891047) for information about the different allelic forms of PspC. |
| NF033838 | PspC_subgroup_1 | 1.17e-04 | 21 | 83 | 623 | 683 | pneumococcal surface protein PspC, choline-binding form. The pneumococcal surface protein PspC, as described in Streptococcus pneumoniae, is a repetitive and highly variable protein, recognized by a conserved N-terminal domain and also by genomic location. This form, subgroup 1, has variable numbers of a choline-binding repeat in the C-terminal region, and is also known as choline-binding protein A. The other form, subgroup 2, is anchored covalently after cleavage by sortase at a C-terminal LPXTG site. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIB28168.1 | 6.40e-136 | 91 | 390 | 2 | 301 |
| QCJ07784.1 | 1.25e-120 | 94 | 386 | 29 | 321 |
| QGH22373.1 | 1.25e-120 | 94 | 386 | 29 | 321 |
| QGH26411.1 | 1.25e-120 | 94 | 386 | 29 | 321 |
| QSX04350.1 | 1.25e-120 | 94 | 386 | 29 | 321 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4U5I_A | 5.88e-16 | 82 | 324 | 64 | 314 | ChainA, Endoglucanase H [Acetivibrio thermocellus ATCC 27405],4U5I_B Chain B, Endoglucanase H [Acetivibrio thermocellus ATCC 27405],4U5K_A Chain A, Endoglucanase H [Acetivibrio thermocellus ATCC 27405],4U5K_B Chain B, Endoglucanase H [Acetivibrio thermocellus ATCC 27405] |
| 4U3A_A | 1.91e-15 | 82 | 323 | 64 | 313 | ChainA, Endoglucanase H [Acetivibrio thermocellus ATCC 27405],4U3A_B Chain B, Endoglucanase H [Acetivibrio thermocellus ATCC 27405] |
| 5BYW_A | 6.74e-14 | 82 | 323 | 64 | 324 | ChainA, Endoglucanase H [Acetivibrio thermocellus ATCC 27405],5BYW_B Chain B, Endoglucanase H [Acetivibrio thermocellus ATCC 27405],5BYW_C Chain C, Endoglucanase H [Acetivibrio thermocellus ATCC 27405],5BYW_D Chain D, Endoglucanase H [Acetivibrio thermocellus ATCC 27405],5BYW_E Chain E, Endoglucanase H [Acetivibrio thermocellus ATCC 27405] |
| 6KDD_A | 8.61e-10 | 99 | 386 | 16 | 317 | endoglucanase[Fervidobacterium pennivorans DSM 9078] |
| 3RJY_A | 1.51e-09 | 94 | 379 | 11 | 313 | CrystalStructure of Hyperthermophilic Endo-beta-1,4-glucanase in complex with substrate [Fervidobacterium nodosum Rt17-B1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P16218 | 2.16e-14 | 82 | 323 | 315 | 564 | Endoglucanase H OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celH PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
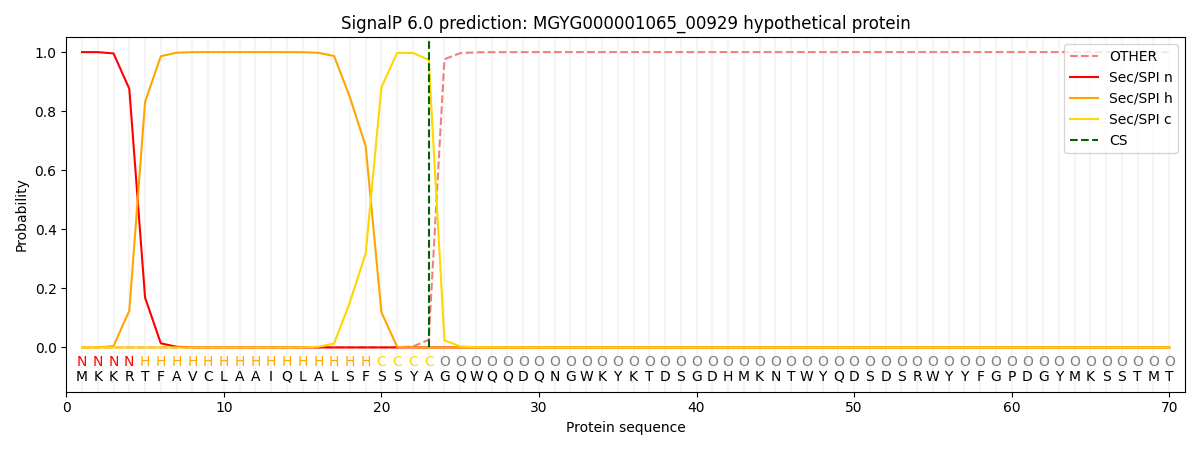
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000418 | 0.998660 | 0.000304 | 0.000229 | 0.000209 | 0.000170 |
