You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001068_01299
You are here: Home > Sequence: MGYG000001068_01299
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Streptococcus sp000187445 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus sp000187445 | |||||||||||
| CAZyme ID | MGYG000001068_01299 | |||||||||||
| CAZy Family | GT113 | |||||||||||
| CAZyme Description | Beta-1,6-galactofuranosyltransferase WbbI | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5040; End: 6092 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT113 | 12 | 349 | 1.8e-102 | 0.9813664596273292 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK09814 | PRK09814 | 1.01e-93 | 1 | 349 | 1 | 329 | sugar transferase. |
| cd03794 | GT4_WbuB-like | 1.34e-05 | 23 | 320 | 56 | 362 | Escherichia coli WbuB and similar proteins. This family is most closely related to the GT1 family of glycosyltransferases. WbuB in E. coli is involved in the biosynthesis of the O26 O-antigen. It has been proposed to function as an N-acetyl-L-fucosamine (L-FucNAc) transferase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUB38817.1 | 1.26e-228 | 1 | 349 | 1 | 349 |
| ALR79904.1 | 2.09e-227 | 1 | 350 | 1 | 350 |
| QEW10543.1 | 1.23e-219 | 1 | 349 | 1 | 349 |
| QKH69719.1 | 7.06e-196 | 1 | 350 | 1 | 350 |
| QBS00018.1 | 1.58e-192 | 1 | 350 | 1 | 350 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3QKW_A | 8.11e-13 | 23 | 341 | 23 | 317 | Structureof Streptococcus parasangunini Gtf3 glycosyltransferase [Streptococcus parasanguinis],3QKW_B Structure of Streptococcus parasangunini Gtf3 glycosyltransferase [Streptococcus parasanguinis],3QKW_C Structure of Streptococcus parasangunini Gtf3 glycosyltransferase [Streptococcus parasanguinis],3QKW_D Structure of Streptococcus parasangunini Gtf3 glycosyltransferase [Streptococcus parasanguinis] |
| 3RHZ_A | 8.49e-13 | 23 | 341 | 31 | 325 | Structureand functional analysis of a new subfamily of glycosyltransferases required for glycosylation of serine-rich streptococcal adhesions [Streptococcus parasanguinis],3RHZ_B Structure and functional analysis of a new subfamily of glycosyltransferases required for glycosylation of serine-rich streptococcal adhesions [Streptococcus parasanguinis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P37749 | 2.82e-42 | 9 | 349 | 3 | 323 | Beta-1,6-galactofuranosyltransferase WbbI OS=Escherichia coli (strain K12) OX=83333 GN=wbbI PE=1 SV=1 |
| Q9AEU1 | 1.52e-18 | 22 | 341 | 20 | 320 | Glucosyltransferase 3 OS=Streptococcus gordonii OX=1302 GN=gtf3 PE=3 SV=1 |
| B3XPQ7 | 1.56e-16 | 10 | 346 | 8 | 325 | Glucosyltransferase 3 OS=Limosilactobacillus reuteri (strain DSM 17509 / CIP 109821 / 100-23) OX=349123 GN=gtf3 PE=3 SV=1 |
| F8KEJ1 | 3.49e-15 | 10 | 346 | 13 | 330 | N-acetylglucosaminyltransferase OS=Limosilactobacillus reuteri (strain ATCC 53608) OX=927703 GN=gtf3 PE=1 SV=1 |
| B5A7L9 | 4.38e-12 | 23 | 341 | 21 | 315 | Glucosyltransferase 3 OS=Streptococcus parasanguinis OX=1318 GN=gtf3 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
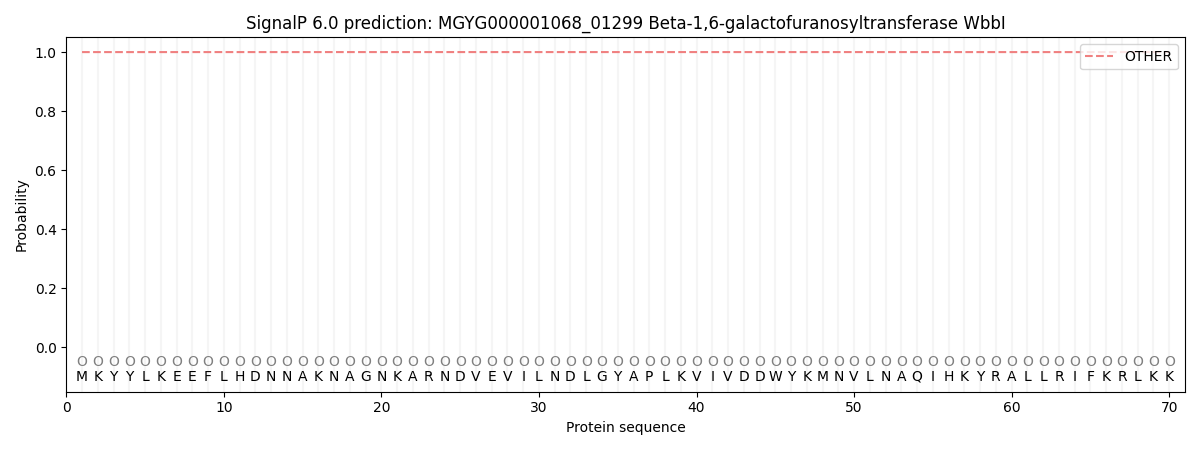
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000055 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
