You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001074_01693
You are here: Home > Sequence: MGYG000001074_01693
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp002251365 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp002251365 | |||||||||||
| CAZyme ID | MGYG000001074_01693 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase F | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2591; End: 3751 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 201 | 355 | 1.3e-28 | 0.8222222222222222 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd13403 | MLTF-like | 9.60e-76 | 193 | 356 | 1 | 160 | membrane-bound lytic murein transglycosylase F (MLTF) and similar proteins. This subfamily includes membrane-bound lytic murein transglycosylase F (MltF, murein lyase F) that degrades murein glycan strands. It is responsible for catalyzing the release of 1,6-anhydromuropeptides from peptidoglycan. Lytic transglycosylase catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do goose-type lysozymes. However, in addition, it also makes a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| PRK10859 | PRK10859 | 5.47e-46 | 37 | 353 | 35 | 448 | membrane-bound lytic murein transglycosylase MltF. |
| COG4623 | MltF | 3.03e-43 | 38 | 357 | 16 | 433 | Membrane-bound lytic murein transglycosylase MltF [Cell wall/membrane/envelope biogenesis, Signal transduction mechanisms]. |
| pfam01464 | SLT | 5.02e-23 | 203 | 308 | 11 | 114 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| cd16896 | LT_Slt70-like | 8.28e-23 | 188 | 317 | 3 | 137 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALO48082.1 | 5.59e-165 | 3 | 386 | 11 | 391 |
| EFC70077.2 | 1.65e-164 | 4 | 386 | 13 | 391 |
| VEH15587.1 | 8.09e-158 | 1 | 386 | 40 | 433 |
| BCS86517.1 | 3.46e-156 | 8 | 386 | 9 | 390 |
| AGB28026.1 | 1.09e-155 | 4 | 386 | 36 | 413 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4OZ9_A | 1.33e-26 | 186 | 353 | 256 | 419 | Crystalstructure of MltF from Pseudomonas aeruginosa complexed with isoleucine [Pseudomonas aeruginosa PAO1] |
| 4OWD_A | 1.44e-26 | 186 | 353 | 263 | 426 | Crystalstructure of MltF from Pseudomonas aeruginosa complexed with cysteine [Pseudomonas aeruginosa PAO1],4OXV_A Crystal structure of MltF from Pseudomonas aeruginosa complexed with valine [Pseudomonas aeruginosa PADK2_CF510],4P0G_A Crystal structure of MltF from Pseudomonas aeruginosa complexed with bulgecin and muropeptide [Pseudomonas aeruginosa PAO1],4P11_A Native crystal structure of MltF Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1] |
| 4OYV_A | 1.44e-26 | 186 | 353 | 263 | 426 | Crystalstructure of MltF from Pseudomonas aeruginosa complexed with leucine [Pseudomonas aeruginosa PAO1] |
| 5AA4_B | 1.68e-26 | 186 | 353 | 249 | 412 | Crystalstructure of MltF from Pseudomonas aeruginosa in complex with cell-wall tetrapeptide [Pseudomonas aeruginosa BWHPSA013],5AA4_D Crystal structure of MltF from Pseudomonas aeruginosa in complex with cell-wall tetrapeptide [Pseudomonas aeruginosa BWHPSA013] |
| 5AA4_A | 1.68e-26 | 186 | 353 | 249 | 412 | Crystalstructure of MltF from Pseudomonas aeruginosa in complex with cell-wall tetrapeptide [Pseudomonas aeruginosa BWHPSA013],5AA4_C Crystal structure of MltF from Pseudomonas aeruginosa in complex with cell-wall tetrapeptide [Pseudomonas aeruginosa BWHPSA013] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A8ZWR8 | 1.14e-30 | 68 | 353 | 155 | 446 | Membrane-bound lytic murein transglycosylase F OS=Desulfococcus oleovorans (strain DSM 6200 / JCM 39069 / Hxd3) OX=96561 GN=mltF PE=3 SV=2 |
| A1U3J1 | 1.73e-29 | 37 | 353 | 32 | 446 | Membrane-bound lytic murein transglycosylase F OS=Marinobacter nauticus (strain ATCC 700491 / DSM 11845 / VT8) OX=351348 GN=mltF PE=3 SV=1 |
| A8AD27 | 1.10e-27 | 41 | 348 | 39 | 444 | Membrane-bound lytic murein transglycosylase F OS=Citrobacter koseri (strain ATCC BAA-895 / CDC 4225-83 / SGSC4696) OX=290338 GN=mltF PE=3 SV=1 |
| Q02RN8 | 5.77e-27 | 91 | 353 | 181 | 450 | Membrane-bound lytic murein transglycosylase F OS=Pseudomonas aeruginosa (strain UCBPP-PA14) OX=208963 GN=mltF PE=3 SV=2 |
| Q8XA45 | 9.75e-27 | 41 | 348 | 39 | 444 | Membrane-bound lytic murein transglycosylase F OS=Escherichia coli O157:H7 OX=83334 GN=mltF PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
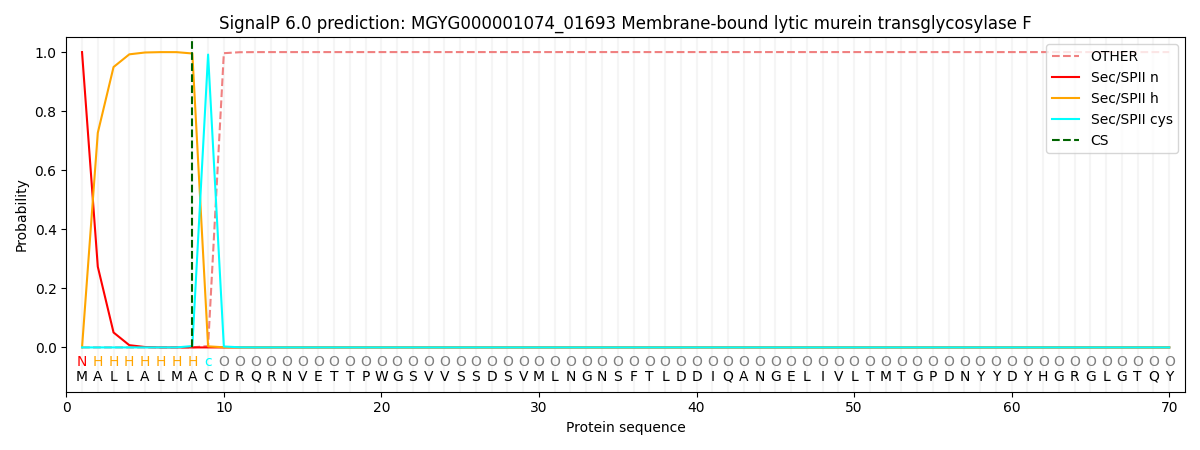
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000007 | 1.000040 | 0.000000 | 0.000000 | 0.000000 |
