You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001159_00253
You are here: Home > Sequence: MGYG000001159_00253
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bifidobacterium pullorum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Bifidobacteriaceae; Bifidobacterium; Bifidobacterium pullorum | |||||||||||
| CAZyme ID | MGYG000001159_00253 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 126875; End: 128161 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 15 | 294 | 2.7e-124 | 0.9964539007092199 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd19608 | GH113_mannanase-like | 0.008 | 4 | 80 | 5 | 81 | Glycoside hydrolase family 113 beta-1,4-mannanase and similar proteins. Mannan endo-1,4-beta mannosidase (E.C 3.2.1.78) randomly cleaves (1->4)-beta-D-mannosidic linkages in mannans, galactomannans and glucomannans and is also called beta-1,4-mannanase, endo-1,4-beta-mannanase, endo-beta-1,4-mannase, beta-mannanase B, beta-1, 4-mannan 4-mannanohydrolase, endo-beta-mannanase, beta-D-mannanase, 1,4-beta-D-mannan mannanohydrolase, and 4-beta-D-mannan mannanohydrolase. (1->4)-beta-linked mannans are polysaccharides with a linear polymer backbone of (1->4)-beta-linked mannose units (in plants and fungi) or alternating mannose and glucose/galactose units (glucomannan in plants and fungi, and galactomannan and galactoglucomannan in plants), such as in the hemicellulose fraction of hard- and softwoods. Complete degradation of mannan requires a series of enzymes, including beta-1,4-mannanase. According to the CAZy database beta-1,4-mannanases are grouped into various glycoside hydrolase (GH) families; GH family 113 beta-1,4-mannanases include mostly bacterial and archaeal sequences. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QAY32595.1 | 1.06e-296 | 1 | 426 | 1 | 426 |
| AIW44813.1 | 6.02e-252 | 1 | 426 | 1 | 426 |
| CBK70291.1 | 6.02e-252 | 1 | 426 | 1 | 426 |
| ADQ02913.1 | 6.02e-252 | 1 | 426 | 1 | 426 |
| QOL47075.1 | 6.02e-252 | 1 | 426 | 1 | 426 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7LR1_A | 5.37e-252 | 1 | 426 | 21 | 446 | ChainA, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR1_B Chain B, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR1_C Chain C, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR1_D Chain D, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR2_A Chain A, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR2_B Chain B, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR2_C Chain C, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR2_D Chain D, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813] |
| 7LR6_A | 4.40e-251 | 1 | 426 | 21 | 446 | ChainA, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR6_B Chain B, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR6_C Chain C, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813],7LR6_D Chain D, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. longum ATCC 55813] |
| 7LQX_A | 5.28e-223 | 2 | 426 | 25 | 449 | ChainA, Glycosyl hydrolase BlGH5_18 [Bifidobacterium longum subsp. infantis ATCC 15697 = JCM 1222 = DSM 20088] |
| 6MP2_A | 1.10e-171 | 1 | 423 | 21 | 445 | Crystalstructure of BlMan5B solved by SIRAS [Bifidobacterium longum DJO10A],6MP2_B Crystal structure of BlMan5B solved by SIRAS [Bifidobacterium longum DJO10A],6MPA_A Chain A, BlMan5B [Bifidobacterium longum DJO10A],6MPA_B Chain B, BlMan5B [Bifidobacterium longum DJO10A] |
| 6MOY_A | 8.96e-171 | 1 | 423 | 21 | 445 | Crystalstructure of the E257A mutant of BlMan5B in complex with GlcNAc (co-crystallization) [Bifidobacterium longum DJO10A],6MOY_B Crystal structure of the E257A mutant of BlMan5B in complex with GlcNAc (co-crystallization) [Bifidobacterium longum DJO10A],6MP7_A Crystal structure of the E257A mutant of BlMan5B in complex with GlcNAc (soaking) [Bifidobacterium longum DJO10A],6MP7_B Crystal structure of the E257A mutant of BlMan5B in complex with GlcNAc (soaking) [Bifidobacterium longum DJO10A],6MPC_A Crystal structure of E257A mutant of BlMan5B [Bifidobacterium longum DJO10A],6MPC_B Crystal structure of E257A mutant of BlMan5B [Bifidobacterium longum DJO10A] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
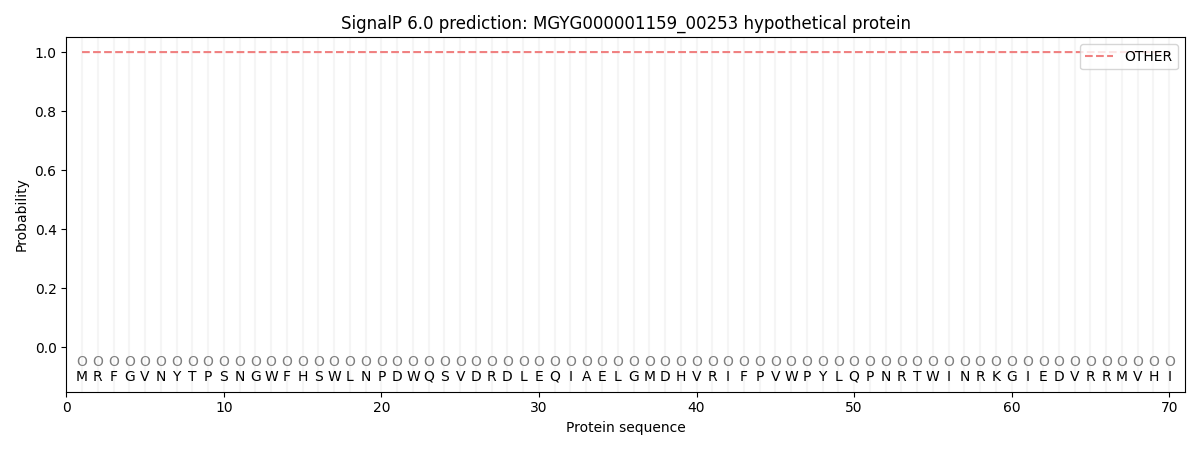
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000061 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
