You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001173_00338
You are here: Home > Sequence: MGYG000001173_00338
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Collinsella sp900759335 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Coriobacteriia; Coriobacteriales; Coriobacteriaceae; Collinsella; Collinsella sp900759335 | |||||||||||
| CAZyme ID | MGYG000001173_00338 | |||||||||||
| CAZy Family | GT0 | |||||||||||
| CAZyme Description | 3-deoxy-manno-octulosonate cytidylyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3652; End: 5301 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd02513 | CMP-NeuAc_Synthase | 1.09e-73 | 7 | 221 | 1 | 220 | CMP-NeuAc_Synthase activates N-acetylneuraminic acid by adding CMP moiety. CMP-N-acetylneuraminic acid synthetase (CMP-NeuAc synthetase) or acylneuraminate cytidylyltransferase catalyzes the transfer the CMP moiety of CTP to the anomeric hydroxyl group of NeuAc in the presence of Mg++. It is the second to last step in the sialylation of the oligosaccharide component of glycoconjugates by providing the activated sugar-nucleotide cytidine 5'-monophosphate N-acetylneuraminic acid (CMP-Neu5Ac), the substrate for sialyltransferases. Eukaryotic CMP-NeuAc synthetases are predominantly located in the nucleus. The activated CMP-Neu5Ac diffuses from the nucleus into the cytoplasm. |
| COG1083 | NeuA | 1.85e-58 | 8 | 228 | 4 | 226 | CMP-N-acetylneuraminic acid synthetase [Cell wall/membrane/envelope biogenesis]. |
| COG3980 | SpsG | 1.19e-30 | 228 | 479 | 2 | 258 | Spore coat polysaccharide biosynthesis protein SpsG, predicted glycosyltransferase [Cell wall/membrane/envelope biogenesis]. |
| pfam02348 | CTP_transf_3 | 1.72e-20 | 9 | 137 | 1 | 121 | Cytidylyltransferase. This family consists of two main Cytidylyltransferase activities: 1) 3-deoxy-manno-octulosonate cytidylyltransferase,, EC:2.7.7.38 catalyzing the reaction:- CTP + 3-deoxy-D-manno-octulosonate <=> diphosphate + CMP-3-deoxy-D-manno-octulosonate, 2) acylneuraminate cytidylyltransferase EC:2.7.7.43, catalyzing the reaction:- CTP + N-acylneuraminate <=> diphosphate + CMP-N-acylneuraminate. NeuAc cytydilyltransferase of Mannheimia haemolytica has been characterized describing kinetics and regulation by substrate charge, energetic charge and amino-sugar demand. |
| cd02517 | CMP-KDO-Synthetase | 3.13e-13 | 7 | 152 | 1 | 135 | CMP-KDO synthetase catalyzes the activation of KDO which is an essential component of the lipopolysaccharide. CMP-KDO Synthetase: 3-Deoxy-D-manno-octulosonate cytidylyltransferase (CMP-KDO synthetase) catalyzes the conversion of CTP and 3-deoxy-D-manno-octulosonate into CMP-3-deoxy-D-manno-octulosonate (CMP-KDO) and pyrophosphate. KDO is an essential component of the lipopolysaccharide found in the outer surface of gram-negative eubacteria. It is also a constituent of the capsular polysaccharides of some gram-negative eubacteria. Its presence in the cell wall polysaccharides of green algae and plant were also discovered. However, they have not been found in yeast and animals. The absence of the enzyme in mammalian cells makes it an attractive target molecule for drug design. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACV55877.1 | 7.31e-239 | 7 | 549 | 6 | 548 |
| API89863.1 | 1.73e-163 | 7 | 545 | 2 | 542 |
| QGS37541.1 | 1.83e-160 | 1 | 547 | 1 | 553 |
| AMB95295.1 | 3.74e-160 | 7 | 545 | 2 | 543 |
| AMB92172.1 | 4.85e-158 | 8 | 545 | 3 | 543 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1QWJ_A | 6.13e-22 | 6 | 226 | 2 | 222 | TheCrystal Structure of Murine CMP-5-N-Acetylneuraminic Acid Synthetase [Mus musculus],1QWJ_B The Crystal Structure of Murine CMP-5-N-Acetylneuraminic Acid Synthetase [Mus musculus],1QWJ_C The Crystal Structure of Murine CMP-5-N-Acetylneuraminic Acid Synthetase [Mus musculus],1QWJ_D The Crystal Structure of Murine CMP-5-N-Acetylneuraminic Acid Synthetase [Mus musculus] |
| 6IFD_A | 3.83e-20 | 9 | 229 | 23 | 249 | CrystalStructure of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae in complex with CDP and Mg2+. [Vibrio cholerae],6IFD_B Crystal Structure of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae in complex with CDP and Mg2+. [Vibrio cholerae],6IFD_C Crystal Structure of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae in complex with CDP and Mg2+. [Vibrio cholerae],6IFD_D Crystal Structure of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae in complex with CDP and Mg2+. [Vibrio cholerae],6IFI_A Crystal Structure of the Apo form of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae [Vibrio cholerae],6IFI_B Crystal Structure of the Apo form of CMP-N-acetylneuraminate Synthetase from Vibrio cholerae [Vibrio cholerae] |
| 6CKJ_A | 2.38e-18 | 9 | 228 | 6 | 227 | N.meningitidis CMP-sialic acid synthetase, ligand-free [Neisseria meningitidis],6CKK_A N. meningitidis CMP-sialic acid synthetase in the presence of CTP and Ca2+ [Neisseria meningitidis],6CKK_B N. meningitidis CMP-sialic acid synthetase in the presence of CTP and Ca2+ [Neisseria meningitidis],6CKL_A N. meningitidis CMP-sialic acid synthetase in the presence of CMP and Neu5Ac2en [Neisseria meningitidis],6CKL_B N. meningitidis CMP-sialic acid synthetase in the presence of CMP and Neu5Ac2en [Neisseria meningitidis],6CKL_C N. meningitidis CMP-sialic acid synthetase in the presence of CMP and Neu5Ac2en [Neisseria meningitidis],6CKM_A N. meningitidis CMP-sialic acid synthetase in the presence of CMP-sialic acid and Ca2+ [Neisseria meningitidis] |
| 1EYR_A | 2.73e-17 | 9 | 228 | 6 | 227 | Structureof a sialic acid activating synthetase, CMP acylneuraminate synthetase in the presence and absence of CDP [Neisseria meningitidis],1EYR_B Structure of a sialic acid activating synthetase, CMP acylneuraminate synthetase in the presence and absence of CDP [Neisseria meningitidis],1EZI_A Structure of a sialic acid activating synthetase, CMP acylneuraminate synthetase in the presence and absence of CDP [Neisseria meningitidis],1EZI_B Structure of a sialic acid activating synthetase, CMP acylneuraminate synthetase in the presence and absence of CDP [Neisseria meningitidis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q45982 | 7.28e-26 | 8 | 229 | 4 | 230 | Post-translational flagellin modification protein B OS=Campylobacter coli OX=195 GN=ptmB PE=3 SV=1 |
| Q58462 | 6.03e-24 | 228 | 479 | 2 | 272 | Uncharacterized protein MJ1062 OS=Methanocaldococcus jannaschii (strain ATCC 43067 / DSM 2661 / JAL-1 / JCM 10045 / NBRC 100440) OX=243232 GN=MJ1062 PE=1 SV=1 |
| Q0P8U6 | 3.03e-22 | 9 | 234 | 4 | 227 | Pseudaminic acid cytidylyltransferase OS=Campylobacter jejuni subsp. jejuni serotype O:2 (strain ATCC 700819 / NCTC 11168) OX=192222 GN=pseF PE=3 SV=1 |
| Q57140 | 9.55e-22 | 9 | 227 | 8 | 227 | Probable N-acylneuraminate cytidylyltransferase OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=neuA PE=3 SV=1 |
| Q0E671 | 1.04e-21 | 2 | 285 | 27 | 311 | N-acylneuraminate cytidylyltransferase A OS=Danio rerio OX=7955 GN=cmasa PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
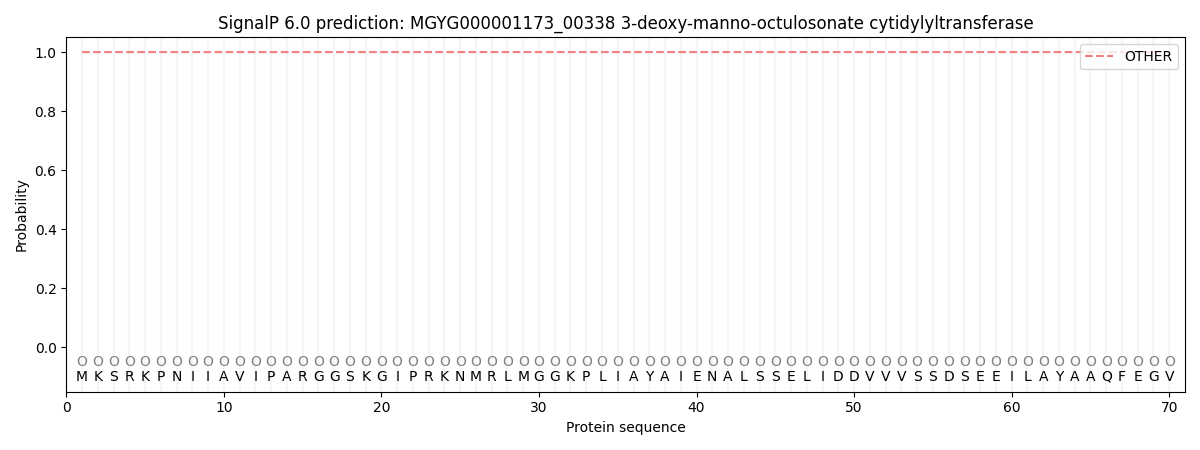
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000041 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
