You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001186_00628
You are here: Home > Sequence: MGYG000001186_00628
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Agathobacter sp900547695 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Agathobacter; Agathobacter sp900547695 | |||||||||||
| CAZyme ID | MGYG000001186_00628 | |||||||||||
| CAZy Family | GH13 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 44661; End: 49784 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH13 | 847 | 1235 | 7.5e-120 | 0.9968354430379747 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd11338 | AmyAc_CMD | 5.75e-163 | 773 | 1270 | 1 | 388 | Alpha amylase catalytic domain found in cyclomaltodextrinases and related proteins. Cyclomaltodextrinase (CDase; EC3.2.1.54), neopullulanase (NPase; EC 3.2.1.135), and maltogenic amylase (MA; EC 3.2.1.133) catalyze the hydrolysis of alpha-(1,4) glycosidic linkages on a number of substrates including cyclomaltodextrins (CDs), pullulan, and starch. These enzymes hydrolyze CDs and starch to maltose and pullulan to panose by cleavage of alpha-1,4 glycosidic bonds whereas alpha-amylases essentially lack activity on CDs and pullulan. They also catalyze transglycosylation of oligosaccharides to the C3-, C4- or C6-hydroxyl groups of various acceptor sugar molecules. Since these proteins are nearly indistinguishable from each other, they are referred to as cyclomaltodextrinases (CMDs). The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| PRK10785 | PRK10785 | 2.13e-85 | 765 | 1342 | 113 | 598 | maltodextrin glucosidase; Provisional |
| cd11316 | AmyAc_bac2_AmyA | 8.04e-52 | 834 | 1269 | 11 | 403 | Alpha amylase catalytic domain found in bacterial Alpha-amylases (also called 1,4-alpha-D-glucan-4-glucanohydrolase). AmyA (EC 3.2.1.1) catalyzes the hydrolysis of alpha-(1,4) glycosidic linkages of glycogen, starch, related polysaccharides, and some oligosaccharides. This group includes Chloroflexi, Dictyoglomi, and Fusobacteria. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| COG0366 | AmyA | 1.74e-51 | 774 | 1309 | 1 | 493 | Glycosidase [Carbohydrate transport and metabolism]. |
| pfam00128 | Alpha-amylase | 3.86e-51 | 847 | 1234 | 1 | 334 | Alpha amylase, catalytic domain. Alpha amylase is classified as family 13 of the glycosyl hydrolases. The structure is an 8 stranded alpha/beta barrel containing the active site, interrupted by a ~70 a.a. calcium-binding domain protruding between beta strand 3 and alpha helix 3, and a carboxyl-terminal Greek key beta-barrel domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QKN24589.1 | 0.0 | 47 | 1348 | 40 | 1343 |
| QKO30210.1 | 0.0 | 47 | 1348 | 40 | 1343 |
| ARP49674.1 | 0.0 | 47 | 1348 | 40 | 1343 |
| VCV21496.1 | 0.0 | 272 | 1376 | 330 | 1441 |
| CBL09766.1 | 0.0 | 273 | 1376 | 331 | 1441 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1JF6_A | 6.25e-75 | 722 | 1350 | 82 | 583 | ChainA, ALPHA AMYLASE II [Thermoactinomyces vulgaris],1JF6_B Chain B, ALPHA AMYLASE II [Thermoactinomyces vulgaris] |
| 1BVZ_A | 6.25e-75 | 722 | 1350 | 82 | 583 | Alpha-amylaseIi (tvaii) From Thermoactinomyces Vulgaris R-47 [Thermoactinomyces vulgaris],1BVZ_B Alpha-amylase Ii (tvaii) From Thermoactinomyces Vulgaris R-47 [Thermoactinomyces vulgaris],1JI2_A Improved X-ray Structure of Thermoactinomyces vulgaris R-47 alpha-Amylase 2 [Thermoactinomyces vulgaris],1JI2_B Improved X-ray Structure of Thermoactinomyces vulgaris R-47 alpha-Amylase 2 [Thermoactinomyces vulgaris],3A6O_A Crystal structure of Thermoactinomyces vulgaris R-47 alpha-amylase 2/acarbose complex [Thermoactinomyces vulgaris],3A6O_B Crystal structure of Thermoactinomyces vulgaris R-47 alpha-amylase 2/acarbose complex [Thermoactinomyces vulgaris] |
| 1JF5_A | 8.46e-75 | 722 | 1350 | 82 | 583 | ChainA, ALPHA AMYLASE II [Thermoactinomyces vulgaris],1JF5_B Chain B, ALPHA AMYLASE II [Thermoactinomyces vulgaris] |
| 1WZM_A | 1.55e-74 | 722 | 1350 | 82 | 583 | ChainA, Alpha-amylase II [Thermoactinomyces vulgaris],1WZM_B Chain B, Alpha-amylase II [Thermoactinomyces vulgaris] |
| 1JIB_A | 2.84e-74 | 722 | 1350 | 82 | 583 | ChainA, NEOPULLULANASE [Thermoactinomyces vulgaris],1JIB_B Chain B, NEOPULLULANASE [Thermoactinomyces vulgaris],1JL8_A Chain A, ALPHA-AMYLASE II [Thermoactinomyces vulgaris],1JL8_B Chain B, ALPHA-AMYLASE II [Thermoactinomyces vulgaris],1VB9_A Chain A, alpha-amylase II [Thermoactinomyces vulgaris],1VB9_B Chain B, alpha-amylase II [Thermoactinomyces vulgaris],2D2O_A Chain A, Neopullulanase 2 [Thermoactinomyces vulgaris],2D2O_B Chain B, Neopullulanase 2 [Thermoactinomyces vulgaris] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P16950 | 3.31e-163 | 459 | 1351 | 37 | 921 | Amylopullulanase OS=Thermoanaerobacter thermohydrosulfuricus OX=1516 GN=apu PE=1 SV=1 |
| P38939 | 1.29e-161 | 459 | 1351 | 37 | 920 | Amylopullulanase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=apu PE=1 SV=2 |
| P36905 | 5.97e-159 | 461 | 1372 | 34 | 938 | Amylopullulanase OS=Thermoanaerobacterium saccharolyticum OX=28896 GN=apu PE=3 SV=2 |
| P38536 | 1.83e-148 | 476 | 1372 | 55 | 937 | Amylopullulanase OS=Thermoanaerobacterium thermosulfurigenes OX=33950 GN=amyB PE=3 SV=2 |
| Q08751 | 3.42e-74 | 722 | 1350 | 82 | 583 | Neopullulanase 2 OS=Thermoactinomyces vulgaris OX=2026 GN=tvaII PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
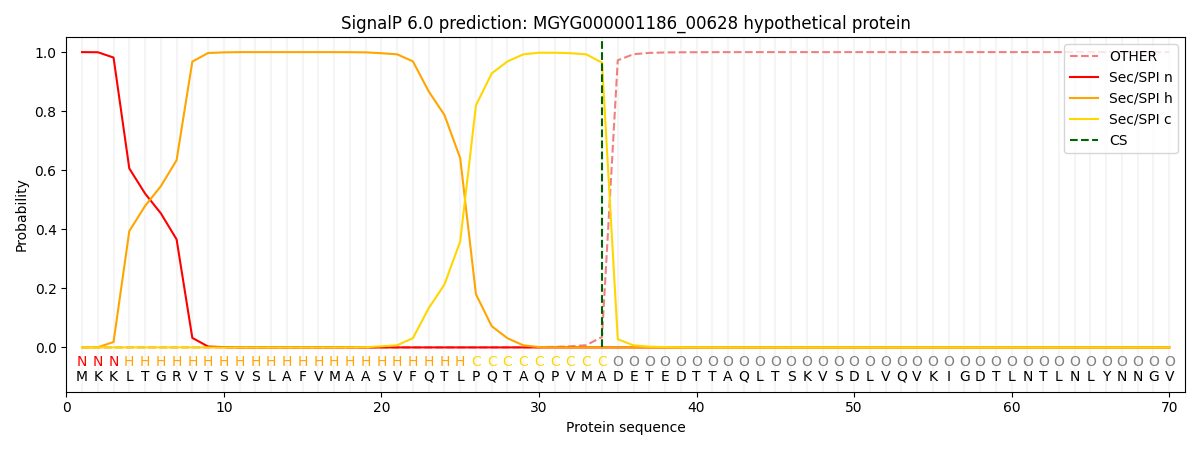
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000245 | 0.999039 | 0.000172 | 0.000186 | 0.000155 | 0.000141 |

